真菌产生多种胞外酶,包括降解多糖的水解酶、降解木质素和开放苯环的氧化酶(Andlar et al. 2018),在养分循环和生态系统稳定方面发挥着重要的作用。木质素和纤维素是影响植物凋落物有机质分解速率的关键因素(Yue et al. 2016),木质素降解主要由漆酶、锰依赖性过氧化物酶、木质素过氧化物酶和多功能过氧化物酶等完成(Andlar et al. 2018),纤维素的降解由内切葡聚糖酶、外切纤维二糖水解酶和β-葡萄糖苷酶等协同完成(Kubicek et al. 2014)。
植物内生真菌是指在生活史的全部或部分阶段定殖在植物内部的一类真菌(Wilson 1995)。内生真菌是当今国际菌物学研究的热点对象(王敏等 2018),不仅具有高度的生物多样性,也具有高度多样化的生态学功能(Rodriguez et al. 2009)。内生真菌不仅定殖于活的健康植物中,还可定殖于植物凋落物中,参与叶和根凋落物中有机质的分解(Guerreiro et al. 2018;Kohout et al. 2021)。
真菌类群是影响凋落物有机质分解的重要因素,如马尾松Pinus massoniana内生散斑壳菌Lophodermium spp.在2个月内造成14%-22%的质量损失,且几乎所有的散斑壳菌都表现出中等或较强的纤维素酶和漆酶(两种最重要的木质纤维素分解酶)阳性反应(Yuan & Chen 2014)。在座囊菌纲Dothideomycetes中,来源于活的和死亡叶片的真菌的纤维素酶活性存在显著差异,但在粪壳菌纲Sordariomycetes中则没有这种差异(U’Ren & Arnold 2016)。格孢腔菌目Pleosporales是座囊菌纲中最大的目,作为附生菌、腐生菌、内生菌或寄生菌、病原菌、真菌或昆虫的重寄生菌和地衣型真菌在世界范围内广泛分布(Hongsanan 2020)。我们前期研究发现格孢腔菌目是禾本科植物的主要优势内生真菌(刘蔚廷等 2019),通过分析格孢腔菌目内生真菌的漆酶和纤维素酶活性,进一步对具有明显漆酶和纤维素酶活性的四绺孢球腔菌科Tetraplosphaeriaceae真菌进行多重对应分析,从而可以探索四绺孢球腔菌科真菌的生态学功能。
1 材料与方法
1.1 序列测定及多基因系统发育树构建
供试菌株为云南省西双版纳纳板河流域国家级自然保护区禾本科植物牛筋草Eleusine indica和疣粒稻Oryza granulata分离的格孢腔菌目内生真菌E00680、E00711、E00595、E00606、E00608、E00610和E00657。采用引物对ITS-1/ITS-4(White et al. 1990)、LR0R/LR5(Vilgalys & Hester 1990)、fRPB2-7cR/ RPB2-5F(Liu et al. 1999)、EF-983F/EF1-2212R(Rehner & Buckley 2002)、bena-T1/bena-T22 (O’Donnell & Cigelnik 1997)扩增核糖体内转录间隔区(internal transcribed spaces and 5.8S rDNA,ITS)、核糖体大亚基(large subunits rDNA,LSU)、RNA聚合酶Ⅱ的第二大亚基(the second largest subunit of RNA polymeraseⅡ,rpb2)、翻译延伸因子-1α基因(translation elongation factor-1 alpha,tef1)和β-微管蛋白基因(beta-tubulin,tub2)。PCR扩增产物由浙江尚亚生物技术有限公司进行测序。采用MAFFT(Katoh & Standley 2013)、BMGE(Dereeper et al. 2010)和SequenceMatrix(Vaidya et al. 2011)对序列进行多重序列比对、校准和连结,用IQTREE(Trifinopoulos et al. 2016)以最大似然法(maximum likelihood,ML)建立系统发育树。
1.2 纤维素和木质素降解能力的测定
羧甲基纤维素钠培养基(g/L):羧甲基纤维素钠(CMC-Na)10,(NH4)2SO4 2.0,K2HPO4 1.0,MgSO4·7H2O 0.5,KCl 0.5,FeSO4·7H2O 0.01,琼脂15.0。菌株在PDA培养基25℃活化培养14d,切取菌饼1块接种于羧甲基纤维素钠培养基平板,25℃暗培养5d,用1mg/mL的刚果红染液覆盖平板染色30min,倒掉刚果红溶液,用蒸馏水清洗,再用1mol/L的NaCl溶液覆盖平板脱色1h,观察记录水解透明圈直径。
愈创木酚培养基:PDA培养基中加入0.04%过滤除菌的愈创木酚。菌株在PDA培养基25℃活化培养14d,切取菌饼1块接种于愈创木酚培养基平板,25℃暗培养5d,观察记录菌落周围红棕色变色圈直径。
1.3 多重对应分析
对四绺孢球腔菌科真菌进行多重对应分析。首先,根据本研究菌株的序列,通过NCBI BLAST搜索从GenBank中检索下载序列,利用R version 4.0.2提取序列、登录号、菌株或克隆号、寄主、分离来源和地理位置等信息,根据多基因系统发育分析确定其分类地位。然后,用SPSS多重对应分析方法探讨四绺孢球腔菌科真菌的寄主、分离来源、地理位置等之间的联系。
2 结果与分析
2.1 系统发育关系
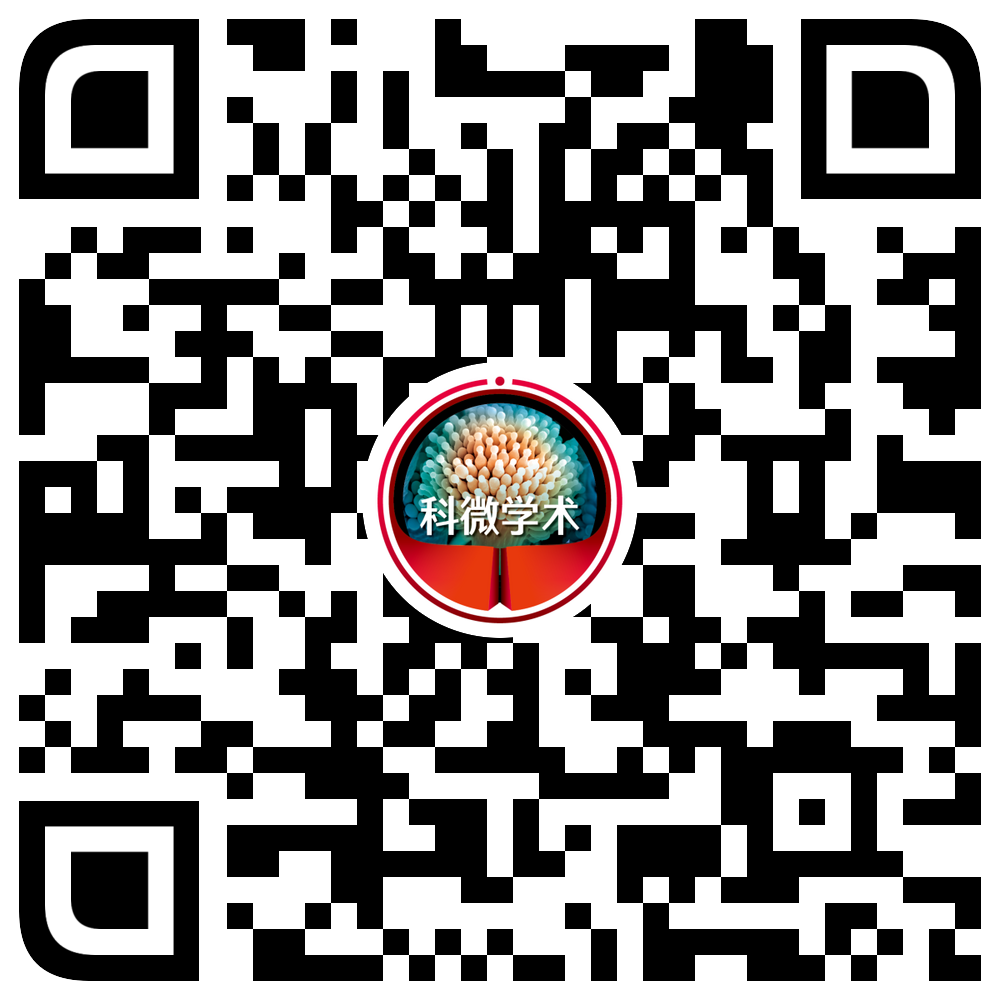
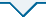
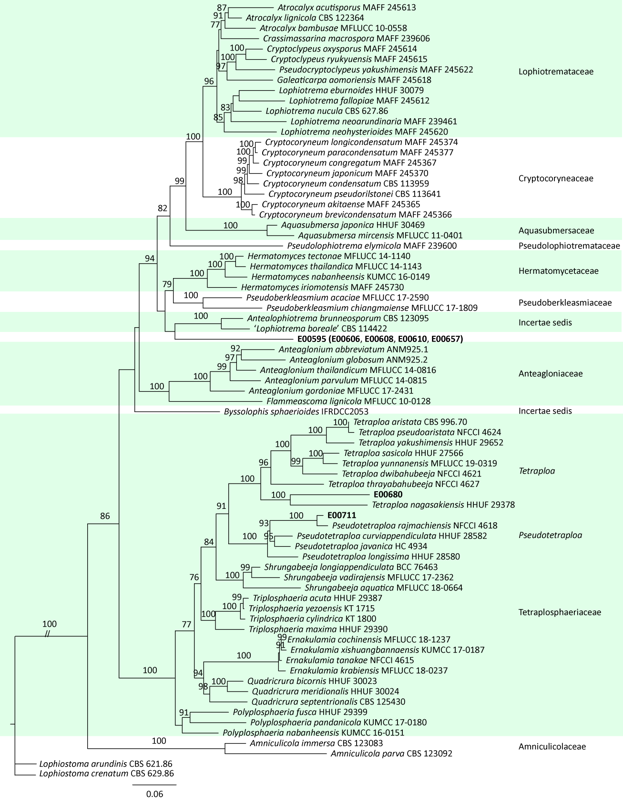
基于ITS、LSU、rpb2、tef1和tub2序列,以格孢腔菌目的扁孔壳菌科Lophiostomataceae的Lophiostoma arundinis CBS 621.86和Lophiostoma crenatum CBS 629.86为外群,构建了本研究供试内生真菌菌株与格孢腔菌目的扁孔皮壳菌科Lophiotremataceae、隐紫胶盘菌科Cryptocoryneaceae、浸水囊菌科Aquasubmersaceae、假扁孔皮壳菌科Pseudolophiotremataceae、肤皮菌科Hermatomycetaceae、假顶丛隔孢霉科Pseudoberkleasmiaceae、前船壳科Anteagloniaceae、四绺孢球腔菌科和近缘的未定属genera incertae sedis代表种的ML树,分支节点处给出≥75%的最大似然自举值。ML树(图1)显示,各科组成的分支最大似然自举值均≥75%。E00595、E00606、E00608、E00610和E00657共5个菌株与未定属的‘Lophiotrema boreale’CBS 114422、Antealophiotrema brunneosporum CBS 123095近缘,但与它们的距离均较远,因此,菌株E00595、E00606、E00608、E00610和E00657应该为不同于‘Lophiotrema boreale’、Antealophiotrema brunneosporum的未知属。菌株E00680和E00711与四绺孢球腔菌科的四绺孢属Tetraploa、假四绺孢属Pseudotetraploa、收缩菌属Shrungabeeja、多重球腔菌属Polyplosphaeria、Ernakulamia、四尾腔菌属Quadricrura和三重球腔菌属Triplosphaeria构建一个最大似然自举值100%的分支,其中菌株E00680与已知的四绺孢属的8个种构成一个最大似然自举值100%的分支,与Tetraploa nagasakiensis近缘但与Tetraploa nagasakiensis距离较远,因此,可能代表了一个与Tetraploa nagasakiensis近缘的未知新种;菌株E00711与已知的假四绺孢属的4个种构成一个最大似然自举值100%的分支,与Pseudotetraploa rajmachiensis近缘但与Pseudotetraploa rajmachiensis距离较远,因此,可能代表了一个与Pseudotetraploa rajmachiensis近缘的未知新种。
图1
图1
四绺孢球腔菌科及近缘真菌的多基因系统发育树
Fig. 1
Multi-loci phylogenetic tree of Tetraplosphaeriaceae and related fungi based on ITS, LSU, rpb2, tef1, tub2, analysed by maximum likelihood (ML).
2.2 纤维素酶和漆酶活性
图2
图2
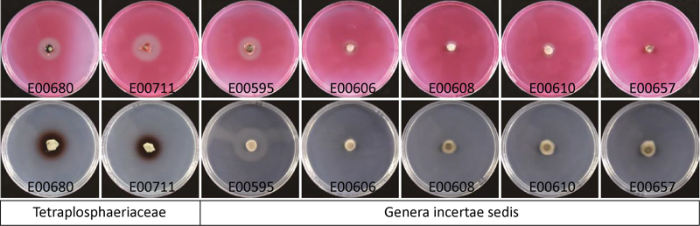
格孢腔菌目2个内生真菌类群在羧甲基纤维素钠培养基和愈创木酚培养基上的菌落
Fig. 2
Colonies of two endophytic fungal taxa of Pleosporales on sodium carboxymethyl cellulose and guaiacol medium.
表1 格孢腔菌目2个内生真菌类群在羧甲基纤维素钠培养基和愈创木酚培养基上的水解透明圈或红棕色变色圈直径
Table 1
| 培养基 Medium | 水解透明圈或红棕色变色圈直径 Diameters of hydrolyzed clear zone or reddish-brown discoloration circle (cm) | ||||||
|---|---|---|---|---|---|---|---|
| Tetraplosphaeriaceae | Genera incertae sedis | ||||||
| E00680 | E00711 | E00595 | E00606 | E00608 | E00610 | E00657 | |
| 羧甲基纤维素钠培养基 Sodium carboxymethyl cellulose medium | 2.09±0.01 c* | 2.59±0.05 a | 2.19±0.03 b | 1.39±0.01 e | 1.16±0.02 f | 1.42±0.03 d | 0 |
| 愈创木酚培养基 Guaiacol medium | 2.69±0.04 a | 2.29±0.03 b | 0 | 0 | 0 | 0 | 0 |
注:a,b,c,d,e,f表示方差分析结果,P<0.05
Note: a, b, c, d, e, f represent the results of anova, P<0.05.
格孢腔菌目的未定属菌株E00595、E00606、E00608、E00610和E00657在羧甲基纤维素钠培养基上培养5d,E00595表现出明显的水解透明圈,水解透明圈直径为2.19cm,E00606、E00608、E00610有较小的水解透明圈,E00657没有水解透明圈;在愈创木酚培养基上培养5d,菌落周围均没有明显的红棕色变色圈。结果表明格孢腔菌目未定属的5株内生真菌仅具有一定的纤维素降解活性。
2.3 四绺孢球腔菌科真菌与寄主、分离来源和地理位置的关联
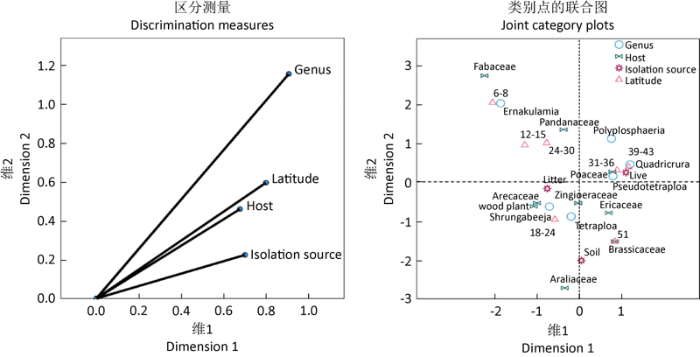
对从GenBank中检索的序列,根据多基因系统发育分析确定属于四绺孢球腔菌科的记录141条,鉴于一些未培养克隆序列信息只有分离来源,因此纳入SPSS多重对应分析的序列66条。基于SPSS多重对应分析的四绺孢球腔菌科真菌的寄主、分离来源、地理位置等的关系(图3),由区分测量图可以看到,四绺孢球腔菌科的属与纬度、寄主和分离来源之间均呈现锐角,说明它们之间都有关联,其中与纬度和寄主的关联最大。由类别点联合图可以看到,分离来源上四绺孢属与土壤、凋落物和活的植物关联较小,地理上与纬度18-24°的区域关联较大,寄主上与棕榈科Arecaceae等木本植物关联稍大,与姜科Zingiberaceae、杜鹃花科Ericaceae、十字花科Brassicaceae和禾本科Poaceae等关联较小,说明四绺孢属主要分布在纬度18-24°N区域的多种寄主植物及其凋落物和土壤;假四绺孢属与禾本科、纬度31-36°N的区域关联,说明假四绺孢属分布在纬度31-36°N区域的禾本科植物上。
图3
图3
基于SPSS多重对应分析的四绺孢球腔菌科真菌的寄主、分离来源和地理分布
Fig. 3
Host, isolated source and geographical distribution of the fungi of Tetraplosphaeriaceae based on SPSS multiple correspondence analysis.
3 讨论
考察生境中不同生态位真菌的相互联系将有助于预测生态系统中物质循环及阐明真菌种群的生态学功能,一些预先定殖于植物中的内生真菌可能是促进凋落物分解的关键因素。对德国林地的山毛榉Fagus sylvatica秋叶和相应凋落物中的真菌种群进行比较,在1年生凋落物中发现了相当比例的叶内生真菌,共现网络分析表明叶片内生真菌是凋落物真菌种群的一个组成部分,是1年生凋落物中最活跃的真菌(Guerreiro et al. 2018)。在极端干旱的纳米比亚纳米布沙海存活着一种多年生禾本科植物Stipagrostis sabulicola,28d的离体分解试验发现20个内生真菌物种中的80%具有降解宿主凋落物的能力,183d的原位田间试验发现有内生真菌的分蘖的分解速度是无菌分蘖的两倍,对凋落物分解过程进行分析发现Stipagrostis sabulicola内生真菌种群中有59%-70%是潜伏腐生菌,这些双生态位真菌在凋落物老化1年后仍占很大比例(58%-62%),利用夜间频繁的非降雨水分事件可以在分蘖衰老后立即启动凋落物的分解,从而最大限度地提高植物束缚的养分在原地循环的可能性,并促进干旱地区普遍存在的营养岛效应 (Wenndt et al. 2021)。
纤维素酶和漆酶是两种最重要的木质纤维素分解酶,不同内生真菌类群的纤维素酶和漆酶活性具有明显差别。在衰亡植物组织分解过程中,前期羧甲基纤维素酶对失重率贡献较大,后期漆酶与过氧化物酶对失重率的贡献升高(陈明蓉等 2020)。在中国北方元宝槭Acer truncatum叶的内生真菌中,常见真菌比稀有真菌具有更高的分解能力,培养16周后所有9种常见真菌造成的质量损失率都超过25%,而12种稀有真菌中只有4种超过25%,推测腐生生活方式可能是常见内生真菌的特征,而稀有内生真菌则因为分解效率较低、产生的繁殖体较少而成为较不常见的内生真菌(Sun et al. 2011)。在马尾松P. massoniana内生真菌中,几乎所有的散斑壳菌都表现出中等或强的纤维素酶和漆酶阳性反应(Yuan & Chen 2014)。分析北美5地的松科、柏科、壳斗科、杜鹃花科、棕榈科、桦木科、杨柳科的18种植物活的叶片中分离的内生真菌及落叶和植物冠层中的死亡叶片分离的落叶真菌和枯叶真菌的木质纤维素降解活性,比较了8个只在活叶中发现的内生真菌OTU和9个只在非活叶中发现的落叶真菌和枯叶真菌OTU的纤维素分解和木质素分解活性,发现所有菌株在体外均未表现出木质素分解活性,而不同种类真菌的纤维素分解活性不同(U’Ren & Arnold 2016)。本研究发现四绺孢球腔菌科的内生真菌E00680和E00711具有明显的纤维素酶和漆酶活性,而格孢腔菌目的一个未定属的内生真菌仅具有较弱的纤维素酶活性、无漆酶活性,表明格孢腔菌目内生真菌的不同类群在木质纤维素降解中存在差异。
四绺孢属是四绺孢球腔菌科的模式属。四绺孢属物种通常腐生在草本植物或腐朽的木头上,也可从土壤中分离到(Tanaka et al. 2009;Hongsanan 2020;Hyde et al. 2020)。本研究的内生真菌菌株E00680分离自云南西双版纳禾本科植物牛筋草根部,系统发育分析显示与Tetraploa nagasakiensis近缘,具有明显的木质纤维素降解能力。已知的假四绺孢属均从禾本科植物中分离,本研究的内生真菌E00711分离自云南西双版纳禾本科植物疣粒稻,系统发育分析显示与Pseudotetraploa rajmachiensis近缘,具有明显的木质纤维素降解能力。多重对应分析发现四绺孢球腔菌科真菌的属与寄主、分离来源和地理位置有关联,但由于四绺孢球腔菌科真菌的记录较少,提供的生态位信息有限,因此多重对应分析提示的与生态位的关联性可能存在偏差。四绺孢属和假四绺孢属两属真菌可在活的健康植物中作为内生真菌存活,并可在植物凋落物和土壤中分离得到,因此我们推测四绺孢属和假四绺孢属两属为内生和腐生双生态位真菌,在植物衰老后可作为一种先锋菌群定殖于植物凋落物中,参与凋落物有机质的分解,并最终进入土壤营腐生生活。进一步开展内生真菌E00680、E00711参与的凋落物离体和原位分解试验以及与群落中其他真菌在凋落物分解中的相互作用将有助于我们深刻认识禾本科植物内生真菌的多样性和生态学功能。
参考文献
Lignocellulose degradation: an overview of fungi and fungal enzymes involved in lignocellulose degradation
DOI:10.1002/elsc.v18.11 URL [本文引用: 2]
Effects of colonization by endophytic fungi of Cunninghamia lanceolata leaves on litter decomposition and associating microbial activities
BLAST-EXPLORER helps you building datasets for phylogenetic analysis
DOI:10.1186/1471-2148-10-8
PMID:20067610
[本文引用: 1]

Background: The right sampling of homologous sequences for phylogenetic or molecular evolution analyses is a crucial step, the quality of which can have a significant impact on the final interpretation of the study. There is no single way for constructing datasets suitable for phylogenetic analysis, because this task intimately depends on the scientific question we want to address, Moreover, database mining softwares such as BLAST which are routinely used for searching homologous sequences are not specifically optimized for this task. Results: To fill this gap, we designed BLAST-Explorer, an original and friendly web-based application that combines a BLAST search with a suite of tools that allows interactive, phylogenetic-oriented exploration of the BLAST results and flexible selection of homologous sequences among the BLAST hits. Once the selection of the BLAST hits is done using BLAST-Explorer, the corresponding sequence can be imported locally for external analysis or passed to the phylogenetic tree reconstruction pipelines available on the Phylogeny.fr platform. Conclusions: BLAST-Explorer provides a simple, intuitive and interactive graphical representation of the BLAST results and allows selection and retrieving of the BLAST hit sequences based a wide range of criterions. Although BLAST-Explorer primarily aims at helping the construction of sequence datasets for further phylogenetic study, it can also be used as a standard BLAST server with enriched output. BLAST-Explorer is available at http://www.phylogeny.fr
Transient leaf endophytes are the most active fungi in 1-year-old beech leaf litter
DOI:10.1007/s13225-017-0390-4 URL [本文引用: 2]
Fungal diversity notes 1151-1276: taxonomic and phylogenetic contributions on genera and species of fungal taxa
DOI:10.1007/s13225-020-00439-5 URL [本文引用: 1]
Refined families of Dothideomycetes: Dothideomycetidae and Pleosporomycetidae
DOI:10.5943/mycosphere URL [本文引用: 2]
MAFFT multiple sequence alignment software version 7: improvements in performance and usability
DOI:10.1093/molbev/mst010 URL [本文引用: 1]
Forest microhabitat affects succession of fungal communities on decomposing fine tree roots
DOI:10.3389/fmicb.2021.541583 URL [本文引用: 1]
Plant cell wall-degrading enzymes and their secretion in plant-pathogenic fungi
DOI:10.1146/phyto.2014.52.issue-1 URL [本文引用: 1]
Phylogenetic relationships among ascomycetes: evidence from an RNA polymerse II subunit
In an effort to establish a suitable alternative to the widely used 18S rRNA system for molecular systematics of fungi, we examined the nuclear gene RPB2, encoding the second largest subunit of RNA polymerase II. Because RPB2 is a single-copy gene of large size with a modest rate of evolutionary change, it provides good phylogenetic resolution of Ascomycota. While the RPB2 and 18S rDNA phylogenies were highly congruent, the RPB2 phylogeny did result in much higher bootstrap support for all the deeper branches within the orders and for several branches between orders of the Ascomycota. There are several strongly supported phylogenetic conclusions. The Ascomycota is composed of three major lineages: Archiascomycetes, Saccharomycetales, and Euascomycetes. Within the Euascomycetes, plectomycetes, and pyrenomycetes are monophyletic groups, and the Pleosporales and Dothideales are distinct sister groups within the Loculoascomycetes. We confirm the placement of Neolecta within the Archiascomycetes, suggesting that fruiting body formation and forcible discharge of ascospores were characters gained early in the evolution of the Ascomycota. These findings show that a slowly evolving protein-coding gene such as RPB2 is useful for diagnosing phylogenetic relationships among fungi.
Diversity of endophytic fungi associated with plants of Poaceae from Yunnan, Zhejiang and Inner Mongolia
Two divergent intragenomic rDNA ITS2 types within a monophyletic lineage of the fungus Fusarium are nonorthologous
DOI:10.1006/mpev.1996.0376 URL [本文引用: 1]
A Beauveria phylogeny inferred from nuclear ITS and EF1-a sequences: evidence for cryptic diversification and links to Cordyceps teleomorphs
Fungal endophytes: diversity and functional roles
DOI:10.1111/j.1469-8137.2009.02773.x
PMID:19236579
[本文引用: 1]

All plants in natural ecosystems appear to be symbiotic with fungal endophytes. This highly diverse group of fungi can have profound impacts on plant communities through increasing fitness by conferring abiotic and biotic stress tolerance, increasing biomass and decreasing water consumption, or decreasing fitness by altering resource allocation. Despite more than 100 yr of research resulting in thousands of journal articles, the ecological significance of these fungi remains poorly characterized. Historically, two endophytic groups (clavicipitaceous (C) and nonclavicipitaceous (NC)) have been discriminated based on phylogeny and life history traits. Here, we show that NC-endophytes represent three distinct functional groups based on host colonization and transmission, in planta biodiversity and fitness benefits conferred to hosts. Using this framework, we contrast the life histories, interactions with hosts and potential roles in plant ecophysiology of C- and NC-endophytes, and highlight several key questions for future work in endophyte biology.
Community composition of endophytic fungi in Acer truncatum and their role in decomposition
DOI:10.1007/s13225-010-0086-5 URL [本文引用: 1]
Molecular taxonomy of bambusicolous fungi: Tetraplosphaeriaceae, a new pleosporalean family with Tetraploa-like anamorphs
DOI:10.3114/sim.2009.64.10
PMID:20169030
[本文引用: 1]

A new pleosporalean family Tetraplosphaeriaceae is established to accommodate five new genera; 1) Tetraplosphaeria with small ascomata and anamorphs belonging to Tetraploa s. str., 2) Triplosphaeria characterised by hemispherical ascomata with rim-like side walls and anamorphs similar to Tetraploa but with three conidial setose appendages, 3) Polyplosphaeria with large ascomata surrounded by brown hyphae and anamorphs producing globose conidia with several setose appendages, 4) Pseudotetraploa, an anamorphic genus, having obpyriform conidia with pseudosepta and four to eight setose appendages, and 5) Quadricrura, an anamorphic genus, having globose conidia with one or two long setose appendages at the apex and four to five short setose appendages at the base. Fifteen new taxa in these genera mostly collected from bamboo are described and illustrated. They are linked by their Tetraploa s. l. anamorphs. To infer phylogenetic placement in the Pleosporales, analyses based on a combined dataset of small- and large-subunit nuclear ribosomal DNA (SSU+LSU nrDNA) was carried out. Tetraplosphaeriaceae, however, is basal to the main pleosporalean clade and therefore its relationship with other existing families was not completely resolved. To evaluate the validity of each taxon and to clarify the phylogenetic relationships within this family, further analyses using sequences from ITS-5.8S nrDNA (ITS), transcription elongation factor 1-alpha (TEF), and beta-tubulin (BT), were also conducted. Monophyly of the family and that of each genus were strongly supported by analyses based on a combined dataset of the three regions (ITS+TEF+BT). Our results also suggest that Tetraplosphaeria (anamorph: Tetraploa s. str.) is an ancestral lineage within this family. Taxonomic placement of the bambusicolous fungi in Astrosphaeriella, Kalmusia, Katumotoa, Massarina, Ophiosphaerella, Phaeosphaeria, Roussoella, Roussoellopsis, and Versicolorisporium, are also discussed based on the SSU+LSU phylogeny.
W-IQ-TREE: a fast online phylogenetic tool for maximum likelihood analysis
DOI:10.1093/nar/gkw256 URL [本文引用: 1]
Diversity, taxonomic composition, and functional aspects of fungal communities in living, senesced, and fallen leaves at five sites across North America
DOI:10.7717/peerj.2768 URL [本文引用: 2]
SequenceMatrix: concatenation software for the fast assembly of multi-gene datasets with character set and codon information
DOI:10.1111/cla.2011.27.issue-2 URL [本文引用: 1]
Rapid genetic identification and mapping of enzymatically amplified ribosomal DNA from several Cryptococcus species
Detailed restriction analyses of many samples often require substantial amounts of time and effort for DNA extraction, restriction digests, Southern blotting, and hybridization. We describe a novel approach that uses the polymerase chain reaction (PCR) for rapid simplified restriction typing and mapping of DNA from many different isolates. DNA fragments up to 2 kilobase pairs in length were efficiently amplified from crude DNA samples of several pathogenic Cryptococcus species, including C. neoformans, C. albidus, C. laurentii, and C. uniguttulatus. Digestion and electrophoresis of the PCR products by using frequent-cutting restriction enzymes produced complex restriction phenotypes (fingerprints) that were often unique for each strain or species. We used the PCR to amplify and analyze restriction pattern variation within three major portions of the ribosomal DNA (rDNA) repeats from these fungi. Detailed mapping of many restriction sites within the rDNA locus was determined by fingerprint analysis of progressively larger PCR fragments sharing a common primer site at one end. As judged by PCR fingerprints, the rDNA of 19 C. neoformans isolates showed no variation for four restriction enzymes that we surveyed. Other Cryptococcus spp. showed varying levels of restriction pattern variation within their rDNAs and were shown to be genetically distinct from C. neoformans. The PCR primers used in this study have also been successfully applied for amplification of rDNAs from other pathogenic and nonpathogenic fungi, including Candida spp., and ought to have wide applicability for clinical detection and other studies.
Analyses of research hotspots and trends of mycology at home and abroad
Why plants harbor complex endophytic fungal communities: insights from perennial bunchgrass Stipagrostis sabulicola in the Namib Sand Sea
DOI:10.3389/fmicb.2021.691584 URL [本文引用: 1]
Amplification and direct sequencing of fungal genes for phylogenetics. In: Innis MA, Gelfand DH, Sninsky JJ, White TJ (eds.) PCR protocols: a guide to methods and applications
Endophyte - the evolution of a term, and clarification of its use and definition
DOI:10.2307/3545919 URL [本文引用: 1]
The role of endophytic fungal individuals and communities in the decomposition of Pinus massoniana needle litter
DOI:10.1371/journal.pone.0105911 URL [本文引用: 2]
Degradation of lignin and cellulose during foliar litter decomposition in an alpine forest river
云南、浙江、内蒙古禾本科植物内生真菌多样性研究