念珠菌病是临床上最常见的真菌感染疾病,从致病情况分析主要分为皮肤黏膜念珠菌病和深部器官念珠菌病(Brown et al. 2012)。近年来,由于抗真菌药物的普遍使用,导致病原菌耐药性不断升高,一些除白念珠菌之外(non-albicans Candida,NAC)物种的临床感染率逐年增加,如光滑念珠菌Candida glabrata、热带念珠菌Candida tropicalis和近平滑念珠菌Candida parapsilosis等。最近的流行病学数据显示,NAC物种占所有系统性念珠菌病比例的35%-65%,其中光滑念珠菌是最常见的NAC物种(Odds 1988;Calderone 2002)。
从全球范围来看,光滑念珠菌血症约占所有念珠菌相关的全身性血液感染的15%(Lim et al. 2012),其患病率因地理区域和个体而异,光滑念珠菌血症在北美洲可高达22%,而在拉丁美洲只占5%左右(Pfaller et al. 2004)。在从尿液分离出的病原真菌中,光滑念珠菌是仅次于白念珠菌的第二常见真菌病原体,由该菌引起的感染可导致外阴阴道念珠菌病(vulvovaginal candidiasis,VVC)。研究数据显示,在从VVC患者病灶部位分离到的NAC中,光滑念珠菌占34.5%(Ozcan et al. 2006);而在从艾滋病患者口腔中分离出的NAC中,光滑念珠菌约14%(Canuto et al. 2000)。与其他NAC相比,由光滑念珠菌引起的感染死亡率更高,据2016年的不完全统计,其感染病死率接近50%(Hull et al. 2000;Thompson et al. 2011;Nishikawa et al. 2016),在骨髓移植患者中,其病死率更是高达100%(Li et al. 2007)。
光滑念珠菌的定植和感染与人群、年龄、地区以及抗菌药的使用情况等因素有关,一般来说免疫功能低下和年老体弱者最易受到光滑念珠菌的感染,但婴幼儿中的光滑念珠菌感染现象极为罕见(Hajjeh et al. 2004)。导致光滑念珠菌易感的因素很多,目前认为HIV感染、老年基础病、静脉留置导管、抗生素等的使用都有可能引起相关的感染性疾病(Silva et al. 2012)。因此对光滑念珠菌生物学功能以及耐药机制的研究是控制并治疗光滑念珠菌感染的基础和前提。
1 光滑念珠菌的生物学研究
1.1 生理生化特性
在沙氏葡萄糖琼脂(SDA)培养基上,光滑念珠菌可以形成光亮且光滑的白色菌落,而在科玛嘉显色培养基(CHROMagar)上通常呈深粉色或紫色(Hospenthal et al. 2006;Daef et al. 2014)。光滑念珠菌细胞(1-4μm)明显小于白念珠菌(4-6μm)、热带念珠菌(4-8μm)和近平滑念珠菌(2.5-4μm)(Calderone 2002)。光滑念珠菌主要以酵母形态生长,很少进行酵母-菌丝形态转换(Glöckner & Cornely 2015)。光滑念珠菌的一般生物学特征见表1。
表1 光滑念珠菌的一般生物学特征
Table 1
| 生物学特性 Biological characteristics | 光滑念珠菌特征 Characteristics of C. glabrata | 参考来源 Reference |
|---|---|---|
| 细胞大小 Cell size | 1-4μm 1-4μm | Calderone 2002 |
| SDA培养基菌落 Colonies on SDA | 光滑/光亮、白色 Smooth/bright, white | Hospenthal et al. 2006; Daef et al. 2014 |
| CHROMagar菌落 Colonies on CHROMagar | 深粉色/紫色 Dark pink/purple | Hospenthal et al. 2006; Daef et al. 2014 |
| 细胞形态 Cell morphology | 酵母,少见有菌丝/假菌丝 Yeast, rarely hyphea/pseudohyphae | Glöckner & Cornely 2015 |
| 细胞倍性 Cell ploidy | 单倍体 Haploid | Wolfe & Shields 1997; Butler et al. 2009 |
| 系统发育分枝 Phylogenetic clade | WGD WGD | Wolfe & Shields 1997; Butler et al. 2009 |
| 碳源利用 Carbon utilization | 可发酵葡萄糖、海藻糖 Ferments glucose and trehalose | Butler et al. 2004 |
| 生物被膜 Biofilm formation | 存在 Present | Kojic & Darouiche 2004; Silva et al. 2009 |
| 主要黏附因子 Major adhesins | Epa蛋白家族 Epa protein family | Kraneveld et al. 2011 |
| 酶分泌 Enzyme secretion | CgYaPsins (CgYps1-11) CgYaPsins (CgYps1-11) | Rasheed et al. 2018 |
光滑念珠菌是单倍体病原菌,在进化树中,与非病原菌酿酒酵母Saccharomyces cerevisiae相近,属于WGD(a whole genome duplication)分支,而白念珠菌和其他念珠菌致病菌属于CTG(translating the CUG codon as seine instead of leucine)分支(Wolfe & Shields 1997;Butler et al. 2009)。比较基因组分析发现,光滑念珠菌在不断进化过程中丢失了许多功能基因,如半乳糖代谢(5个基因)、磷酸盐代谢(4个基因)、防御与毒性相关(3个基因)以及氮硫代谢(3个基因)等,导致某些代谢途径完全丧失(Dujon et al. 2004),如不能利用半乳糖和一些工程改造的氨基酸,仅能发酵和吸收葡萄糖和海藻糖(Butler et al. 2004)。虽然光滑念珠菌的交配途径与酿酒酵母保持同源,且在临床上经常分离出两种不同交配类型菌株,但由于其单倍体特性,且交配相关基因存在缺陷(Silva et al. 2011;Brunke & Hube 2013),尚未发现其存在典型的交配现象。此外,光滑念珠菌自身不能合成烟酸、吡哆醇和硫胺素(Colombo et al. 2007),在缺乏这几种营养的环境中无法生存,通常由哺乳动物等宿主环境补偿。
1.2 基因组稳定性
临床上分离的光滑念珠菌的基因组非常不稳定,经常发生不规则的变化如节段性复制、染色体重排等(Poláková et al. 2009)。丹麦的一项研究表明,通过对200多株临床光滑念珠菌的分析,发现至少通过两种不同的机制形成“新”染色体,即染色体间易位和节段性复制(Ahmad et al. 2013)。这些染色体带有重复的基因,如在耐药性中起作用的ABC转运蛋白(ATP-binding cassette transporter)家族基因、与表面黏附有关的细胞壁相关基因等,可能在病原菌与宿主相互作用中发挥一定作用(Poláková et al. 2009)。值得注意的是,即使在系统发育上相近的光滑念珠菌分离株也具有不同的染色体重排现象,表明该病原菌的基因组很不稳定。此外,光滑念珠菌基因组不稳定和功能适应性改变不仅仅在侵染宿主期间发生,也可以在短时间内且没有极端选择压力的实验室条件下发生(Ahmad et al. 2014),这很有可能与光滑念珠菌在适应性进化中获得的习性有关。
1.3 生物被膜
约65%-80%的人类感染疾病与病原菌生物被膜的形成有关(Davies 2003)。光滑念珠菌可以黏附在宿主组织和非生物表面如导尿管、血管导管和起搏器上形成多层生物被膜结构(Kojic & Darouiche 2004;Silva et al. 2009),它们对抗真菌药物具有高度的耐药性(Donlan & Costerton 2002)。生物被膜的形成始于细胞的黏附,之后细胞分裂(增殖)最终形成细胞外基质(成熟),细胞表面的相互作用是由细胞壁中的特殊蛋白介导的,这些蛋白称为黏附蛋白。由于光滑念珠菌酵母-菌丝转换效率极低(Glöckner & Cornely 2015),黏附作用以及生物被膜是该菌尤为重要的毒力因子。
黏附是生物被膜形成的第一步。光滑念珠菌可以表达大量的编码黏附相关的基因,与相近物种如无致病性的酿酒酵母进行基因组比较,发现黏附蛋白编码基因的数量与致病性之间存在相关性(Timmermans et al. 2018)。光滑念珠菌中约有44个黏附基因位于染色体的端粒附近,极易发生重排或者非等位基因同源重组(de Groot et al. 2008),这会导致黏附蛋白产生变异从而改善对特定基质的黏附,进而增加了光滑念珠菌的致病性及对不同环境条件的适应性。
与Als蛋白在白念珠菌生物被膜形成中的作用类似,Epa(epithelial adhesion)蛋白家族是光滑念珠菌黏附蛋白中最重要的一类。其中EPA1主要在生物被膜细胞指数生长期中表达(Kraneveld et al. 2011),也被证实是光滑念珠菌体外黏附上皮细胞的主要黏附蛋白,同时参与人巨噬细胞样细胞的黏附(Cormack et al. 1999)。而EPA6和EPA7主要在生物被膜形成或即将形成的过程中表达(Kraneveld et al. 2011),人工突变EPA6基因会使光滑念珠菌的生物被膜降低约3倍(Iraqui et al. 2005)。EPA基因的表达受到多种因素调控,其中Yak1激酶和Swi/Snf染色质重构复合物可以缓解调控端粒沉默对EPA6和EPA7基因表达的抑制,从而使编码的黏附蛋白在生物被膜形成中发挥重要作用(d’Enfert & Janbon 2016)。研究表明,Epa蛋白可能与人体细胞中的碳水化合物相互作用,如Epa1和Epa7可以与肾脏或者大脑组织结构的N-聚糖互作(Ielasi et al. 2016),而Epa6可以结合到纤连蛋白等(Satala et al. 2016)。有趣的是,葡萄糖感应相关基因突变会影响生物被膜的形成。此外,尿路环境由于富含烟酸,会促进光滑念珠菌EPA1、EPA6、EPA7黏附基因的表达(Domergue et al. 2005),这可能会增加该菌在尿路中的定植。
光滑念珠菌生物被膜通常是集群的细胞紧密排列形成一个多层的结构,没有菌丝或者假菌丝。成熟的光滑念珠菌生物被膜厚度大约是白念珠菌生物被膜的一半(75-90μm)(Kucharíková et al. 2015),具有较高的蛋白质和碳水化合物(Silva et al. 2010)。细胞密度是生物被膜抗真菌药物的重要因素,通过对唑类和两性霉素B类等药物的物理隔离从而产生一种可诱导的耐药,基本不涉及遗传基因的变化(Ramage et al. 2012)。
光滑念珠菌可以通过形成生物被膜来逃避宿主的免疫机制、抵抗抗真菌药物胁迫以及承受来自于其他微生物的竞争压力,是其高感染率高致死率的重要毒力因子。生物被膜往往是抗菌治疗后复发的一个重要原因,因此开发抗真菌药物以防止或者彻底抑制生物被膜的形成是一项艰巨的挑战。
1.4 酶分泌
除了黏附和形成生物被膜,念珠菌还能释放水解酶来破坏宿主组织从而发挥其毒力,其中包括蛋白酶、磷脂酶、脂肪酶和溶血素等。与其他念珠菌不同,光滑念珠菌不产生蛋白酶(Cormack et al. 1999)。但光滑念珠菌可以产生其他酶类,如磷脂酶,可以将磷脂水解成脂肪酸,破坏宿主粘膜,从而有效入侵组织(das Mohan & Ballal 2008);也可以产生溶血素,用以降解宿主的血红蛋白从而获取铁元素来供自己生长(Luo & Samaranayake 2002),但是人们对光滑念珠菌溶血活性的基因表达了解甚少。目前也有研究证实,有11种假定的糖基磷脂酰肌醇(GPI)锚定的天冬氨酰蛋白酶家族,称为CgYapsins(CgYps1-11),也是光滑念珠菌重要的毒性因子(Rasheed et al. 2018)。
1.5 形态转换
表型可塑性对念珠菌快速适应外界环境非常重要。研究发现,光滑念珠菌在含有CuSO4的培养基上可以进行一个自发可逆的高频形态转换系统。主要包括3种菌落表型:白色(WH)、浅棕色(LB)和深棕色(DB)(Lachke et al. 2000)。这主要是由于不同菌落细胞中的MTII基因(编码金属硫蛋白II)转录速率不同,从而影响将CuSO4还原为CuS的能力,因此显示出不同的颜色(Mehra et al. 1989;Mehra et al. 1990)。有研究指出,在CuSO4培养基上表现为DB表型的菌落具有更强的毒性(Srikantha et al. 2008)。
由此可见,光滑念珠菌的形态转换代表了一种分级系统,这种分级系统在一定程度上促进了光滑念珠菌毒性的研究。
1.6 其他侵袭力
念珠菌在附着后通常会侵入人体上皮细胞层,对于白念珠菌来说,其侵袭方式主要有两种,一种是宿主细胞诱导的内吞作用,另外一种是通过菌丝的主动渗透(Wächtler et al. 2011),其中后者在水解酶以及其他因素协同下,在入侵深层组织和器官中起着至关重要的作用。
那么光滑念珠菌又是如何在没有菌丝的情况下进入宿主系统并引起感染性疾病呢?研究表明,光滑念珠菌入侵的可能途径是通过手术创伤或一些医源性原因破坏了人体的自然屏障(Perlroth et al. 2007),也有可能通过形成假菌丝进行入侵(Csank & Haynes 2000)。在体内,它可能主要依靠内吞作用,且几乎不会损害宿主细胞(Westwater et al. 2007)。相较白念珠菌,光滑念珠菌入侵几乎不诱导炎性因子,也不刺激嗜中性粒细胞,所以没有较强的促炎性细胞因子反应(Westwater et al. 2007;Jacobsen et al. 2010)。
2 耐药性
2.1 耐药现状
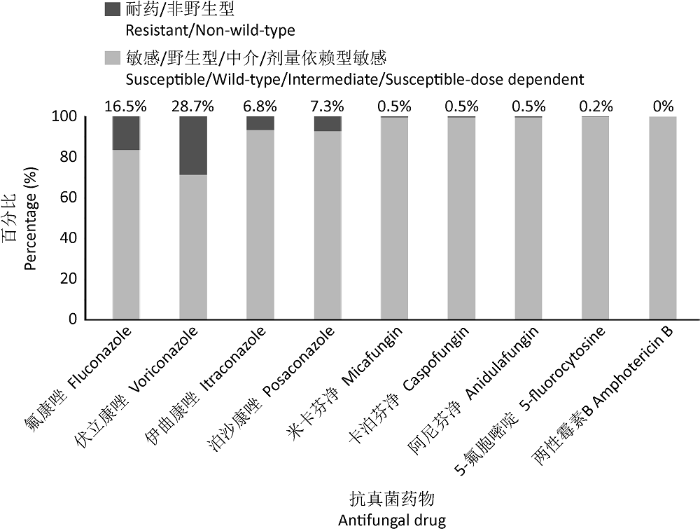
由于抗真菌剂的大量使用,临床上分离到的NAC越来越多,且NAC对氟康唑的耐药性呈不断上升趋势。在丹麦的一项监测研究中,发现2012-2015年与2008-2011年和2004-2007年相比,唑类药物耐药增加,敏感度分别为60.6%、65.2%和68.5%,其中光滑念珠菌耐药占据了主导地位(Astvad et al. 2018)。世界范围内的耐药趋势表明,白念珠菌的氟康唑耐药率为0.3%,热带念珠菌为3.2%,近平滑念珠菌为3.9%,而光滑念珠菌高达8.1%。其中光滑念珠菌耐药率最高的为北美(10.6%),其次为亚太地区(6.8%)、欧洲(4.9%)和拉丁美洲(3.7%)(Pfaller et al. 2019),光滑念珠菌在中国(2009-2014)对多种抗真菌药物的耐药比例见图1(Hou et al. 2017)。
图1
图1
光滑念珠菌耐药菌株所占比例(中国,2009-2014)
Fig. 1
The proportion of resistant strains of Candida glabrata (China, 2009-2014).
棘白霉素药物是治疗系统性感染念珠菌病的首选药物,它不同于唑类和多烯类易引发耐药性,是极有价值的抗真菌药物。从历史上看,光滑念珠菌是对棘白霉素类抗性最强的念珠菌,其耐药率(1.7%-3.5%)远高于白念珠菌(0-0.1%)、近平滑念珠菌(0-0.1%)和热带念珠菌(0.5%-0.7%)(Pfaller et al. 2019)。由于棘白霉素类作为一线药物被广泛使用,这种耐药性的出现令人担忧。
2.2 耐药机制
研究发现光滑念珠菌在经过氟康唑治疗后能迅速产生耐药性,在接触过氟康唑的菌株中,大约20%可出现耐药现象(Krcmery & Barnes 2002)。这种耐药性的出现与下述几种机制有关:(1)与药物外排蛋白有关,其中ATP结合转运蛋白研究得最为广泛,如Cdr1(candida drug resistance 1)和Mdr1(multi-drug resistance 1)等,上述基因的功能获得性突变会使药物外排蛋白高表达(Gohar et al. 2017),Pdr1(pleomorphic drug resistance homolog 1)也参与上述基因的表达,PDR1的功能突变可引起下游的药物外排泵基因表达上调,从而使得光滑念珠菌对唑类药物产生耐受性(Vale-Silva et al. 2016);(2)与药物靶标和甾醇稳态有关,如Upc2A调控包括ERG11基因在内的甾醇生物合成途径中酶的表达,唑类药物以此为药物靶点进而抑制麦角固醇的合成,最终破坏细胞结构。而该类基因突变会使其与唑类药物的亲和性大大降低,从而产生耐药性(Eliopoulos et al. 2002);(3)与光滑念珠菌毒性有关,如前所述光滑念珠菌形成的生物被膜对抗真菌药物具有耐药性,此外,Pdr1突变会使光滑念珠菌增强对宿主上皮细胞的黏附,从而降低巨噬细胞的吞噬,使得唑类药物药效降低(Vale-Silva et al. 2013);(4)与基因组不稳定有关,在耐药菌落中存在极少部分的小突变体(petite mutants),这些突变体内部分耐药相关基因如CDR1会上调,从而对唑类药物耐药(Sanglard et al. 2001)。
随着对唑类药物敏感性降低,棘白霉素类药物已经成为对抗光滑念珠菌的一线治疗药物。在所有棘白霉素耐药菌株中有71%属于光滑念珠菌,它们均具有突变的FKS基因(Pfaller et al. 2019)。FKS基因负责编码葡聚糖合成酶的催化亚基,基因突变通常发生在FKS1或FKS2中,可使棘白霉素的药效降低50-3 000倍(Hendrickson et al. 2019)。目前为止,FKS突变是唯一已知的导致棘白霉素耐药性的基因。
多烯类药物如两性霉素B虽长期以来被视为重要的抗真菌药物,但因其可导致患者肾毒性而被限制使用。其靶点为麦角固醇,光滑念珠菌多烯类药物耐药性的产生与ERG2、ERG6或ERG11的基因突变有关(Vandeputte et al. 2008;Hull et al. 2012a;Hull et al. 2012b)。最近研究发现,与DNA修复相关的MSH2基因突变与耐药突变(PDR1、ERG6、FKS1、FKS2)的增加有关,不同程度上导致了念珠菌对氟康唑、两性霉素B、棘白霉素类耐药(Healey et al. 2016)。
2.3 抗真菌药物的研发
尽管棘白霉素类药物的总体耐药率很低,但是美国一些机构报告称,与唑类药物的耐药性一样,光滑念珠菌对棘白霉素的耐药性也会随着对这类药物的接触使用而不断上升(Malani et al. 2005),这就迫切需要寻找能够针对光滑念珠菌感染的治疗新策略。
新型抗真菌药物在克服耐药性和避免与现有药物相关的副作用方面,可能更具优势,如VT-1129/1161/1598(特异性抑制Cyp51,也称羊毛甾醇-14-α-脱甲基酶)、CD101和SCY-078(抑制葡聚糖合成酶)、T-2307(破坏真菌线粒体膜)和F901318(抑制真菌嘧啶生物合成)等(Wiederhold 2017),但目前仍处于临床前分析评估阶段。
3 讨论
侵袭性非白念珠菌感染率逐年升高,而随着抗真菌药物在危重病人中频繁使用、常规的预防性使用,以及抗真菌类杀虫剂的广泛使用,抗真菌耐药问题会日益显著。事实证明,由于光滑念珠菌天然的药物耐受性,唑类和多烯类药物已经不能很好地治疗相关念珠菌病,而现有棘白霉素类药物的临床治疗也需严格防范其耐药趋势。因此现阶段探索光滑念珠菌的毒力因子以及致病机理有助于新药的研制开发,同时也要规范临床用药,必要时采取联合用药。总的来说,只有更加全面地了解光滑念珠菌的流行病学、致病机理和耐药机制,才能对其临床感染进行有效地防控。
参考文献
Small chromosomes among Danish Candida glabrata isolates originated through different mechanisms
Genome structure and dynamics of the yeast pathogen Candida glabrata
Update from a 12-year nationwide fungemia surveillance: increasing intrinsic and acquired resistance causes concern
Hidden killers: human fungal infections
Two unlike cousins: Candida albicans and C. glabrata infection strategies
DOI:10.1111/cmi.12091
URL
PMID:23253282
[本文引用: 1]

Candida albicans and C. glabrata are the two most common pathogenic yeasts of humans, yet they are phylogenetically, genetically and phenotypically very different. In this review, we compare and contrast the strategies of C. albicans and C. glabrata to attach to and invade into the host, obtain nutrients and evade the host immune response. Although their strategies share some basic concepts, they differ greatly in their outcome. While C. albicans follows an aggressive strategy to subvert the host response and to obtain nutrients for its survival, C. glabrata seems to have evolved a strategy which is based on stealth, evasion and persistence, without causing severe damage in murine models. However, both fungi are successful as commensals and as pathogens of humans. Understanding these strategies will help in finding novel ways to fight Candida, and fungal infections in general.
Evolution of the MAT locus and its Ho endonuclease in yeast species
Evolution of pathogenicity and sexual reproduction in eight Candida genomes
DOI:10.1038/nature08064
URL
PMID:19465905
[本文引用: 3]

Candida species are the most common cause of opportunistic fungal infection worldwide. Here we report the genome sequences of six Candida species and compare these and related pathogens and non-pathogens. There are significant expansions of cell wall, secreted and transporter gene families in pathogenic species, suggesting adaptations associated with virulence. Large genomic tracts are homozygous in three diploid species, possibly resulting from recent recombination events. Surprisingly, key components of the mating and meiosis pathways are missing from several species. These include major differences at the mating-type loci (MTL); Lodderomyces elongisporus lacks MTL, and components of the a1/2 cell identity determinant were lost in other species, raising questions about how mating and cell types are controlled. Analysis of the CUG leucine-to-serine genetic-code change reveals that 99% of ancestral CUG codons were erased and new ones arose elsewhere. Lastly, we revise the Candida albicans gene catalogue, identifying many new genes.
Introduction and historical perspectives
Determinants for the development of oropharyngeal colonization or infection by fluconazole-resistant Candida strains in HIV-infected patients
Prospective observational study of candidemia in Sao Paulo, Brazil: incidence rate, epidemiology, and predictors of mortality
An adhesin of the yeast pathogen Candida glabrata mediating adherence to human epithelial cells
Candida glabrata displays pseudohyphal growth
DOI:10.1111/j.1574-6968.2000.tb09216.x
URL
PMID:10913876
[本文引用: 1]

The ability to undergo morphological change has been reported as an advantageous trait in fungal pathogenesis. Here we demonstrate that Candida glabrata ATCC2001, like diploid Saccharomyces cerevisiae strains, forms elongated chains of pseudohyphal cells on solid nitrogen starvation media (SLAD). Constrictions were apparent between adjoining cells; no parallel-sided hyphae were seen and pseudohyphae invaded the agar. When SLAD was supplemented with ammonium sulfate both C. glabrata and diploid S. cerevisiae strains lost their ability to undergo pseudohyphal growth. However, on this media C. glabrata yeast cells invaded the agar in a similar fashion to the invasive growth mode exhibited by haploid strains of S. cerevisiae cultured on rich media such as YPD. C. glabrata was not capable of invading YPD demonstrating that the process of filamentation is distinct in these two fungi. To our knowledge this is the first report to demonstrate that C. glabrata can undergo morphological change and grow as an invasive filamentous organism.
Evaluation of chromogenic media and seminested PCR in the identification of Candida species
DOI:10.1590/S1517-83822014005000040
URL
PMID:24948942
[本文引用: 3]

Identification of Candida cultured from various clinical specimens to the species level is increasingly necessary for clinical laboratories. Although sn PCR identifies the species within hours but its cost-effectiveness is to be considered. So there is always a need for media which help in the isolation and identification at the species level. The study aimed to evaluate the performance of different chromogenic media and to compare the effectiveness of the traditional phenotypic methods vs. seminested polymerase chain reaction (sn PCR) for identification of Candida species. One hundred and twenty seven Candida strains isolated from various clinical specimens were identified by conventional methods, four different chromogenic media and sn PCR. HiCrome Candida Differential and CHROMagar Candida media showed comparably high sensitivities and specificities in the identification of C. albicans, C. tropicalis, C. glabrata and C. krusei. CHROMagar Candida had an extra advantage of identifying all C. parapsilosis isolates. CHROMagar-Pal's medium identified C. albicans, C. tropicalis and C. krusei with high sensitivities and specificities, but couldn't identify C. glabrata or C. parapsilosis. It was the only medium that identified C. dubliniensis with a sensitivity and specificity of 100%. Biggy agar showed the least sensitivities and specificities. The overall concordance of the snPCR compared to the conventional tests including CHROMAgar Candida in the identification of Candida species was 97.5%. The use of CHROMAgar Candida medium is an easy and accurate method for presumptive identification of the most commonly encountered Candida spp.
Proteinase and phospholipase activity as virulence factors in Candida species isolated from blood
DOI:10.1016/s1130-1406(08)70050-0
URL
PMID:19071887
[本文引用: 1]

The number of nosocomial blood stream infections due to Candida species has increased over the past few decades. In order to establish an infection, opportunistic pathogens have to evade the immune system, survive, divide in the host environment, and spread to other tissues. Proteinase and phospholipase secretion has been implicated as potential virulence factors for some Candida species responsible for catheter related candidemia in intensive care unit (ICU) patients with indwelling devices. We therefore have aimed at demonstrating the secretion of proteinase and phospholipase enzymes as virulent factors by Candida species isolated from blood samples collected from ICUs, dialysis units and oncology units. One hundred and fourteen isolates of Candida species were obtained from the blood samples and the isolates include 37 Candida albicans, 7 Candida glabrata, 5 Candida guilliermondii, 3 Candida kefyr, 45 Candida krusei, 5 Candida parapsilosis, and 12 Candida tropicalis. Proteinase assay was performed by using the Staib et al method. Phospholipase assay was performed by using the method of Samaranayake et al. Precipitation zone (Pz value) was determined. The percentage of isolates which produced detectable amounts of proteinase is 74.56% and 44.73% of isolates produced detectable amounts of phospholipase. We believe that production of both phospholipase and proteinase enzimes could be an important virulence factor for several Candida species.
Understanding biofilm resistance to antibacterial agents
DOI:10.1038/nrd1008 URL PMID:12563302 [本文引用: 1]
The cell wall of the human pathogen Candida glabrata: differential incorporation of novel adhesin-like wall proteins
Biofilm formation in Candida glabrata: what have we learnt from functional genomics approaches? FEMS Yeast Research
Nicotinic acid limitation regulates silencing of Candida adhesins during UTI
DOI:10.1126/science.1108640
URL
PMID:15774723
[本文引用: 1]

The adherence of Candida glabrata to host cells is mediated, at least in part, by the EPA genes, a family of adhesins encoded at subtelomeric loci, where they are subject to transcriptional silencing. We show that normally silent EPA genes are expressed during murine urinary tract infection (UTI) and that the inducing signal is the limitation of nicotinic acid (NA), a precursor of nicotinamide adenine dinucleotide (NAD+). C. glabrata is an NA auxotroph, and NA-induced EPA expression is likely the result of a reduction in NAD+ availability for the NAD+-dependent histone deacetylase Sir2p. The adaptation of C. glabrata to the host, therefore, involves a loss of metabolic capacity and exploitation of the resulting auxotrophy to signal a particular host environment.
Biofilms: survival mechanisms of clinically relevant microorganisms
DOI:10.1128/cmr.15.2.167-193.2002
URL
PMID:11932229
[本文引用: 1]

Though biofilms were first described by Antonie van Leeuwenhoek, the theory describing the biofilm process was not developed until 1978. We now understand that biofilms are universal, occurring in aquatic and industrial water systems as well as a large number of environments and medical devices relevant for public health. Using tools such as the scanning electron microscope and, more recently, the confocal laser scanning microscope, biofilm researchers now understand that biofilms are not unstructured, homogeneous deposits of cells and accumulated slime, but complex communities of surface-associated cells enclosed in a polymer matrix containing open water channels. Further studies have shown that the biofilm phenotype can be described in terms of the genes expressed by biofilm-associated cells. Microorganisms growing in a biofilm are highly resistant to antimicrobial agents by one or more mechanisms. Biofilm-associated microorganisms have been shown to be associated with several human diseases, such as native valve endocarditis and cystic fibrosis, and to colonize a wide variety of medical devices. Though epidemiologic evidence points to biofilms as a source of several infectious diseases, the exact mechanisms by which biofilm-associated microorganisms elicit disease are poorly understood. Detachment of cells or cell aggregates, production of endotoxin, increased resistance to the host immune system, and provision of a niche for the generation of resistant organisms are all biofilm processes which could initiate the disease process. Effective strategies to prevent or control biofilms on medical devices must take into consideration the unique and tenacious nature of biofilms. Current intervention strategies are designed to prevent initial device colonization, minimize microbial cell attachment to the device, penetrate the biofilm matrix and kill the associated cells, or remove the device from the patient. In the future, treatments may be based on inhibition of genes involved in cell attachment and biofilm formation.
Genome evolution in yeasts
DOI:10.1038/nature02579
URL
PMID:15229592
[本文引用: 1]

Identifying the mechanisms of eukaryotic genome evolution by comparative genomics is often complicated by the multiplicity of events that have taken place throughout the history of individual lineages, leaving only distorted and superimposed traces in the genome of each living organism. The hemiascomycete yeasts, with their compact genomes, similar lifestyle and distinct sexual and physiological properties, provide a unique opportunity to explore such mechanisms. We present here the complete, assembled genome sequences of four yeast species, selected to represent a broad evolutionary range within a single eukaryotic phylum, that after analysis proved to be molecularly as diverse as the entire phylum of chordates. A total of approximately 24,200 novel genes were identified, the translation products of which were classified together with Saccharomyces cerevisiae proteins into about 4,700 families, forming the basis for interspecific comparisons. Analysis of chromosome maps and genome redundancies reveal that the different yeast lineages have evolved through a marked interplay between several distinct molecular mechanisms, including tandem gene repeat formation, segmental duplication, a massive genome duplication and extensive gene loss.
Antifungal resistance in pathogenic fungi
DOI:10.1086/344058
URL
PMID:12384841
[本文引用: 1]

Pathogenic fungi are the cause of life-threatening infections in an increasing number of immunocompromised patients. The intrinsic resistance to antifungal therapy observed in some genera, along with the development of resistance during treatment in others, is becoming a major problem in the management of these diseases. We reviewed the epidemiology of the most common systemic fungal infections for which antifungal resistance is a potential problem, the mechanisms of antifungal resistance, the correlation between in vitro susceptibility testing and clinical outcome, and the clinical implications of antifungal resistance.
Candida glabrata-unique features and challenges in the clinical management of invasive infections
Expression patterns of ABC transporter genes in fluconazole-resistant Candida glabrata
DOI:10.1007/s11046-016-0074-8
URL
PMID:27744635
[本文引用: 1]

Clinical management of fungal diseases is compromised by the emergence of antifungal drug resistance in fungi, which leads to elimination of available drug classes as treatment options. An understanding of antifungal resistance at molecular level is, therefore, essential for the development of strategies to combat the resistance. This study presents the assessment of molecular mechanisms associated with fluconazole resistance in clinical Candida glabrata isolates originated from Iran. Taking seven distinct fluconazole-resistant C. glabrata isolates, real-time PCRs were performed to evaluate the alternations in the regulation of the genes involved in drug efflux including CgCDR1, CgCDR2, CgSNQ2, and CgERG11. Gain-of-function (GOF) mutations in CgPDR1 alleles were determined by DNA sequencing. Cross-resistance to fluconazole, itraconazole, and voriconazole was observed in 2.5 % of the isolates. In the present study, six amino acid substitutions were identified in CgPdr1, among which W297R, T588A, and F575L were previously reported, whereas D243N, H576Y, and P915R are novel. CgCDR1 overexpression was observed in 57.1 % of resistant isolates. However, CgCDR2 was not co-expressed with CgCDR1. CgSNQ2 was upregulated in 71.4 % of the cases. CgERG11 overexpression does not seem to be associated with azole resistance, except for isolates that exhibited azole cross-resistance. The pattern of efflux pump gene upregulation was associated with GOF mutations observed in CgPDR1. These results showed that drug efflux mediated by adenosine-5-triphosphate (ATP)-binding cassette transporters, especially CgSNQ2 and CgCDR1, is the predominant mechanism of fluconazole resistance in Iranian isolates of C. glabrata. Since some novel GOF mutations were found here, this study also calls for research aimed at investigating other new GOF mutations to reveal the comprehensive understanding about efflux-mediated resistance to azole antifungal agents.
Incidence of bloodstream infections due to Candida species and in vitro susceptibilities of isolates collected from 1998 to 2000 in a population-based active surveillance program
DOI:10.1128/jcm.42.4.1519-1527.2004
URL
PMID:15070998
[本文引用: 1]

To determine the incidence of Candida bloodstream infections (BSI) and antifungal drug resistance, population-based active laboratory surveillance was conducted from October 1998 through September 2000 in two areas of the United States (Baltimore, Md., and the state of Connecticut; combined population, 4.7 million). A total of 1,143 cases were detected, for an average adjusted annual incidence of 10 per 100,000 population or 1.5 per 10,000 hospital days. In 28% of patients, Candida BSI developed prior to or on the day of admission; only 36% of patients were in an intensive care unit at the time of diagnosis. No fewer than 78% of patients had a central catheter in place at the time of diagnosis, and 50% had undergone surgery within the previous 3 months. Candida albicans comprised 45% of the isolates, followed by C. glabrata (24%), C. parapsilosis (13%), and C. tropicalis (12%). Only 1.2% of C. albicans isolates were resistant to fluconazole (MIC, > or = 64 microg/ml), compared to 7% of C. glabrata isolates and 6% of C. tropicalis isolates. Only 0.9% of C. albicans isolates were resistant to itraconazole (MIC, > or = 1 micro g/ml), compared to 19.5% of C. glabrata isolates and 6% of C. tropicalis isolates. Only 4.3% of C. albicans isolates were resistant to flucytosine (MIC, > or = 32 microg/ml), compared to < 1% of C. parapsilosis and C. tropicalis isolates and no C. glabrata isolates. As determined by E-test, the MICs of amphotericin B were > or = 0.38 microg/ml for 10% of Candida isolates, > or =1 microg/ml for 1.7% of isolates, and > or = 2 microg/ml for 0.4% of isolates. Our findings highlight changes in the epidemiology of Candida BSI in the 1990s and provide a basis upon which to conduct further studies of selected high-risk subpopulations.
Prevalent mutator genotype identified in fungal pathogen Candida glabrata promotes multi-drug resistance
Antifungal resistance: a concerning trend for the present and future.
Presumptive identification of Candida species other than C. albicans, C. krusei, and C. tropicalis with the chromogenic medium CHROMagar Candida
Molecular epidemiology and antifungal susceptibility of Candida glabrata in China (August 2009 to July 2014): a multi-center study
DOI:10.3389/fmicb.2017.00880
URL
PMID:28588560
[本文引用: 1]

Candida glabrata is an increasingly important cause of invasive candidiasis. In China, relatively little is known of the molecular epidemiology of C. glabrata and of its antifungal susceptibility patterns. Here we studied 411 non-duplicate C. glabrata isolates from 411 patients at 11 hospitals participating in the National China Hospital Invasive Fungal Surveillance Net program (CHIF-NET; 2010-2014). Genotyping was performed using multilocus sequence typing (MLST) employing six genetic loci and by microsatellite analysis. Antifungal susceptibility testing was performed using Sensititre YeastOne YO10 methodology. Of 411 isolates, 35 sequence types (ST) were identified by MLST and 79 different genotypes by microsatellite typing; the latter had higher discriminatory power than MLST in the molecular typing of C. glabrata. Using MLST, ST7 and ST3 were the most common STs (66.4 and 9.5% of all isolates, respectively) with 24 novel STs identified; the most common microsatellite types were T25 (30.4% of all isolates) and T31 (12.4%). Resistance to fluconazole (MIC > 32 mug/mL) was seen in 16.5% (68/411) of isolates whilst MICs of >0.5 mug/mL for voriconazole, >2 mug/mL for itraconazole and >2 mug/mL for posaconazole were seen for 28.7, 6.8, and 7.3% of isolates, respectively; 14.8% of all isolates cross-resistant/non-wide-type to fluconazole and voriconazole. Fluconazole resistant rates increased 3-fold over the 5-year period whilst that of isolates with non-WT MICs to voriconazole, 7-fold. All echinocandins exhibited >99% susceptibility rates against all isolates but notably one isolate exhibited multi-drug resistance to the azoles and echinocandins. The study has provided a global picture of the molecular epidemiology and drug resistance rates of C. glabrata in China during the period of the study.
Two clinical isolates of Candida glabrata exhibiting reduced sensitivity to amphotericin B both harbor mutations in ERG2
Facultative sterol uptake in an ergosterol-deficient clinical isolate of Candida glabrata harboring a missense mutation in ERG11 and exhibiting cross-resistance to azoles and amphotericin B
DOI:10.1128/AAC.06253-11
URL
PMID:22615281
[本文引用: 1]

We identified a clinical isolate of Candida glabrata (CG156) exhibiting flocculent growth and cross-resistance to fluconazole (FLC), voriconazole (VRC), and amphotericin B (AMB), with MICs of >256, >256, and 32 mug ml(-1), respectively. Sterol analysis using gas chromatography-mass spectrometry (GC-MS) revealed that CG156 was a sterol 14alpha-demethylase (Erg11p) mutant, wherein 14alpha-methylated intermediates (lanosterol was >80% of the total) were the only detectable sterols. ERG11 sequencing indicated that CG156 harbored a single-amino-acid substitution (G315D) which nullified the function of native Erg11p. In heterologous expression studies using a doxycycline-regulatable Saccharomyces cerevisiae erg11 strain, wild-type C. glabrata Erg11p fully complemented the function of S. cerevisiae sterol 14alpha-demethylase, restoring growth and ergosterol synthesis in recombinant yeast; mutated CG156 Erg11p did not. CG156 was culturable using sterol-free, glucose-containing yeast minimal medium ((glc)YM). However, when grown on sterol-supplemented (glc)YM (with ergosta 7,22-dienol, ergosterol, cholestanol, cholesterol, Delta(7)-cholestenol, or desmosterol), CG156 cultures exhibited shorter lag phases, reached higher cell densities, and showed alterations in cellular sterol composition. Unlike comparator isolates (harboring wild-type ERG11) that became less sensitive to FLC and VRC when cultured on sterol-supplemented (glc)YM, facultative sterol uptake by CG156 did not affect its azole-resistant phenotype. Conversely, CG156 grown using (glc)YM with ergosterol (or with ergosta 7,22-dienol) showed increased sensitivity to AMB; CG156 grown using (glc)YM with cholesterol (or with cholestanol) became more resistant (MICs of 2 and >64 mug AMB ml(-1), respectively). Our results provide insights into the consequences of sterol uptake and metabolism on growth and antifungal resistance in C. glabrata.
Evidence for mating of the “asexual” yeast Candida albicans in a mammalian host
Lectin-glycan interaction network-based identification of host receptors of microbial pathogenic adhesins
DOI:10.1128/mBio.00584-16
URL
PMID:27406561
[本文引用: 1]

UNLABELLED: The first step in the infection of humans by microbial pathogens is their adherence to host tissue cells, which is frequently based on the binding of carbohydrate-binding proteins (lectin-like adhesins) to human cell receptors that expose glycans. In only a few cases have the human receptors of pathogenic adhesins been described. A novel strategy-based on the construction of a lectin-glycan interaction (LGI) network-to identify the potential human binding receptors for pathogenic adhesins with lectin activity was developed. The new approach is based on linking glycan array screening results of these adhesins to a human glycoprotein database via the construction of an LGI network. This strategy was used to detect human receptors for virulent Escherichia coli (FimH adhesin), and the fungal pathogens Candida albicans (Als1p and Als3p adhesins) and C. glabrata (Epa1, Epa6, and Epa7 adhesins), which cause candidiasis. This LGI network strategy allows the profiling of potential adhesin binding receptors in the host with prioritization, based on experimental binding data, of the most relevant interactions. New potential targets for the selected adhesins were predicted and experimentally confirmed. This methodology was also used to predict lectin interactions with envelope glycoproteins of human-pathogenic viruses. It was shown that this strategy was successful in revealing that the FimH adhesin has anti-HIV activity. IMPORTANCE: Microbial pathogens may express a wide range of carbohydrate-specific adhesion proteins that mediate adherence to host tissues. Pathogen attachment to host cells is achieved through the binding of these lectin-like adhesins to glycans on human glycoproteins. In only a few cases have the human receptors of pathogenic adhesins been described. We developed a new strategy to predict these interacting receptors. Therefore, we developed a novel LGI network that would allow the mapping of potential adhesin binding receptors in the host with prioritization, based on the experimental binding data, of the most relevant interactions. New potential targets for the selected adhesins (bacterial uroepithelial FimH from E. coli and fungal Epa and Als adhesins from C. glabrata and C. albicans) were predicted and experimentally confirmed. This methodology was also used to predict lectin interactions with human-pathogenic viruses and to discover whether FimH adhesin has anti-HIV activity.
The Yak1p kinase controls expression of adhesins and biofilm formation in Candida glabrata in a Sir4p‐dependent pathway
DOI:10.1111/j.1365-2958.2004.04475.x
URL
PMID:15686569
[本文引用: 1]

Biofilm is the predominant type of microbial development in natural environments, and potentially represents a major form of resistance or source of recurrence during host infection. Although a large number of studies have focussed on the genetics of bacterial biofilm formation, very little is known about the genes involved in this type of growth in fungi. A genetic screen for Candida glabrata Biofilm mutants was performed using a 96-well plate model of biofilm formation. Study of the isolated mutant strains allowed the identification of four genes involved in biofilm formation (RIF1, SIR4, EPA6 and YAK1). Epa6p is a newly identified adhesin required for biofilm formation in this pathogenic yeast. EPA6 and its close paralogue EPA7 are located in subtelomeric regions and their transcription is regulated by Sir4p and Rif1p, two proteins involved in subtelomeric silencing. Biofilm growth conditions induce the transcription of EPA6 and EPA7: this is dependent on the presence of an intact subtelomeric silencing machinery and is independent of the Mpk1p signalling pathway. Finally, the kinase Yak1p is required for expression of both adhesin genes and acts through a subtelomeric silencing machinery-dependent pathway.
Candida glabrata persistence in mice does not depend on host immunosuppression and is unaffected by fungal amino acid auxotrophy
DOI:10.1128/IAI.01244-09
URL
PMID:20008535
[本文引用: 1]

Candida glabrata has emerged as an important fungal pathogen of humans, causing life-threatening infections in immunocompromised patients. In contrast, mice do not develop disease upon systemic challenge, even with high infection doses. In this study we show that leukopenia, but not treatment with corticosteroids, leads to fungal burdens that are transiently increased over those in immunocompetent mice. However, even immunocompetent mice were not capable of clearing infections within 4 weeks. Tissue damage and immune responses to microabscesses were mild as monitored by clinical parameters, including blood enzyme levels, histology, myeloperoxidase, and cytokine levels. Furthermore, we investigated the suitability of amino acid auxotrophic C. glabrata strains for in vitro and in vivo studies of fitness and/or virulence. Histidine, leucine, or tryptophan auxotrophy, as well as a combination of these auxotrophies, did not influence in vitro growth in rich medium. The survival of all auxotrophic strains in immunocompetent mice was similar to that of the parental wild-type strain during the first week of infection and was only mildly reduced 4 weeks after infection, suggesting that C. glabrata is capable of utilizing a broad range of host-derived nutrients during infection. These data suggest that C. glabrata histidine, leucine, or tryptophan auxotrophic strains are suitable for the generation of knockout mutants for in vivo studies. Notably, our work indicates that C. glabrata has successfully developed immune evasion strategies enabling it to survive, disseminate, and persist within mammalian hosts.
Identification and differential gene expression of adhesin-like wall proteins in Candida glabrata biofilms
DOI:10.1007/s11046-011-9446-2
URL
PMID:21769633
[本文引用: 3]

An important initial step in biofilm development and subsequent establishment of fungal infections by the human pathogen Candida glabrata is adherence to a surface. Adherence is mediated through a large number of differentially regulated cell wall-bound adhesins. The fungus can modify the incorporation of adhesins in the cell wall allowing crucial adaptations to new environments. In this study, expression and cell wall incorporation of C. glabrata adhesins were evaluated in biofilms cultured in two different media: YPD and a semi-defined medium SdmYg. Tandem mass spectrometry of isolated C. glabrata cell walls identified 22 proteins including six adhesins: the novel adhesins Awp5 and Awp6, Epa3 and the previously identified adhesins Epa6, Awp2 and Awp4. Regulation of expression of these and other relevant adhesin genes was investigated using real-time qPCR analysis. For most adhesin genes, significant up-regulation was observed in biofilms in at least one of the culturing media. However, this was not the case for EPA6 and AWP2, which is consistent with their gene products already being abundantly present in planktonic cultures grown in YPD medium. Furthermore, most of the adhesin genes tested also show medium-dependent differential regulation. These results underline the idea that many adhesins in C. glabrata are involved in biofilm formation and that their expression is tightly regulated and dependent on environmental conditions and growth phase. This may contribute to its potential to form resilient biofilms and cause infection in various host tissues.
Non-albicans Candida spp. causing fungaemia: pathogenicity and antifungal resistance
DOI:10.1053/jhin.2001.1151
URL
PMID:12014897
[本文引用: 1]

Non-albicans Candida (NAC) species cause 35-65% of all candidaemias in the general patient population. They occur more frequently in cancer patients, mainly in those with haematological malignancies and bone marrow transplant (BMT) recipients (40-70%), but are less common among intensive care unit (ITU) and surgical patients (35-55%), children (1-35%) or HIV-positive patients (0-33%). The proportion of NAC species among Candida species is increasing: over the two decades to 1990, NAC represented 10-40% of all candidaemias. In contrast, in 1991-1998, they represented 35-65% of all candidaemias. The most common NAC species are C. parapsilosis (20-40% of all Candida species), C. tropicalis (10-30%), C. krusei (10-35%) and C. glabrata (5-40%). Although these four are the most common, at least two other species are emerging: C. lusitaniae causing 2-8% of infections, and C. guilliermondii causing 1-5%. Other NAC species, such as C. rugosa, C. kefyr, C. stellatoidea, C. norvegensis and C. famata are rare, accounting for less than 1% of fungaemias in man. In terms of virulence and pathogenicity, some NAC species appear to be of lower virulence in animal models, yet behave with equal or greater virulence in man, when comparison is made with C. albicans. Mortality due to NAC species is similar to C. albicans, ranging from 15% to 35%. However, there are differences in both overall and attributable mortality among species: the lowest mortality is associated with C. parapsilosis, the highest with C. tropicalis and C. glabrata (40-70%). Other NAC species including C. krusei are associated with similar overall mortality to C. albicans (20-40%). Mortality in NAC species appears to be highest in ITU and surgical patients, and somewhat lower in cancer patients, children and HIV-positive patients. There is no difference between overall and attributable mortality, with the exception of C. glabrata which tends to infect immunocompromised individuals. While the crude mortality is low, attributable mortality (fungaemia-associated mortality) is higher than with C. albicans. There are several specific risk factors for particular NAC species: C. parapsilosis is related to foreign body insertion, neonates and hyperalimentation; C. krusei to azole prophylaxis and along with C. tropicalis to neutropenia and BMT; C. glabrata to azole prophylaxis, surgery and urinary or vascular catheters; C. lusitaniae and C. guilliermondii to previous polyene (amphotericin B or nystatin) use; and C. rugosa to burns. Antifungal susceptibility varies significantly in contrast to C. albicans: some NAC species are inherently or secondarily resistant to fluconazole; for example, 75% of C. krusei isolates, 35% of C. glabrata, 10-25% of C. tropicalis and C. lusitaniae. Amphotericin B resistance is also seen in a small proportion: 5-20% of C. lusitaniae and C. rugosa, 10-15% of C. krusei and 5-10% of C. guilliermondii. Other NAC species are akin to C. albicans-susceptible to both azoles and polyenes (C. parapsilosis, the majority of C. guilliermondii strains and C. tropicalis). Therefore, 'species directed' therapy should be administered for fungaemia according to the species identified-amphotericin B for C. krusei and C. glabrata, fluconazole for other species, including polyene-resistant or tolerant Candida species (C. lusitaniae, C. guilliermondii). In vitro susceptibility testing should be performed for most species of NAC in addition to removal of any foreign body to optimize management.
Candida infections of medical devices
DOI:10.1128/cmr.17.2.255-267.2004
URL
PMID:15084500
[本文引用: 2]

The number of indwelling medical devices is escalating, and an increasing proportion of device-related infections are being caused by Candida spp. Candida spp. produce biofilms on synthetic materials, which facilitates adhesion of the organisms to devices and renders them relatively refractory to medical therapy. Management of device-related Candida infections can be challenging. Removal of the infected device is generally needed to establish cure of Candida infections of medical devices. However, since the pathogenesis of Candida bloodstream infection is complicated, more studies are necessary to determine the role of catheter exchange in patients with both gastrointestinal tract mucositis and indwelling catheters. The medical and economic impact of these infections is enormous.
In vivo Candida glabrata biofilm development on foreign bodies in a rat subcutaneous model
DOI:10.1093/jac/dku447
URL
PMID:25406296
[本文引用: 1]

OBJECTIVES: Biofilm studies have been mostly dedicated to the major human fungal pathogen Candida albicans, whereas much less is known about this virulence factor in Candida glabrata, certainly under in vivo conditions. This study provides a deeper understanding of the biofilm development of C. glabrata, its architecture and susceptibility profile to fluconazole and echinocandins. METHODS: In vitro and in vivo C. glabrata biofilms were developed inside serum-coated triple-lumen catheters placed in 24-well polystyrene plates or implanted subcutaneously in the back of a rat, respectively. Scanning electron microscopy and confocal scanning laser microscopy were used to visualize the biofilm architecture. Quantitative real-time PCR was used to demonstrate the expression profile of EPA1, EPA3, EPA6 and AWP1-AWP7 during in vivo biofilm formation. RESULTS: Mature biofilms were observed within the first 48 h and the amount of biofilm reached its maximum by 6 days. Architecturally, mature C. glabrata biofilms consisted of a thick network of yeast cells embedded in an extracellular matrix. Moreover, in vivo biofilms were susceptible to echinocandin drugs, whereas fluconazole remained ineffective. Gene expression profiling revealed that EPA3, EPA6, AWP2, AWP3 and AWP5 were up-regulated in in vivo biofilms compared with in vitro biofilms. CONCLUSIONS: C. glabrata is a unique microorganism, which, despite the lack of transition to the hyphal form, formed thick biofilms inside foreign bodies in vivo. To our knowledge, this is the first study that has described in vivo C. glabrata biofilm development and its architectural changes in detail and provides an insight into the susceptibility profile, as well as the gene expression machinery, of biofilm-associated infections.
Phenotypic switching in Candida glabrata involves phase-specific regulation of the Metallothionein gene MT-IIand the newly discovered hemolysin gene HLP
Candida glabrata, an emerging oral opportunistic pathogen
DOI:10.1177/154405910708600304
URL
PMID:17314251
[本文引用: 1]

Following the widespread use of immunosuppressive therapy and broad-spectrum antimycotic prophylaxis, C. glabrata has emerged as an important opportunistic pathogen in the oral mucosa. In the past, studies on the virulence factors and host-pathogen interactions of this organism were scarce, but continued to rise in recent years. Denture-wearing, immunosuppression, antibiotic therapy, and aging are risk factors for oral colonization or infection with C. glabrata. Compared with C. albicans, C. glabrata exhibits lower oral keratinocyte-adherence capacity, but higher denture-surface-adherence ability. The role of extracellular hydrolase production in the virulence of this organism does not appear to be as important as it is in C. albicans pathogenesis. Although traditionally thought of as a non-transforming yeast organism, both phenotypic switching and pseudohyphal formation have recently been identified in C. glabrata, but their role in pathogenesis is not known. With the exception of granulocyte monocyte colony-stimulating factor, C. glabrata triggers a lower proinflammatory cytokine response in oral epithelial cells than does C. albicans, in a strain-dependent manner. C. glabrata is less susceptible to killing by human beta-defensins than is C. albicans and exhibits various degrees of resistance to the antifungal activity of salivary histatins and mucins. In addition, C. glabrata possesses both innate and acquired resistance against antifungal drugs, due to its ability to modify ergosterol biosynthesis, mitochondrial function, or antifungal efflux. This resistance allows for its relative overgrowth over other susceptible species and may contribute to the recent emergence of C. glabrata infections in chronically immunocompromised populations. Further investigations on the virulence and host-pathogen interactions of C. glabrata are needed to better define the pathogenesis of oral C. glabrata infection in susceptible hosts.
Candida and invasive candidiasis: back to basics
Candida glabrata, an emerging fungal pathogen, exhibits superior relative cell surface hydrophobicity and adhesion to denture acrylic surfaces compared with Candida albicans
DOI:10.1034/j.1600-0463.2002.1100902.x URL [本文引用: 1]
Candida glabrata fungemia: experience in a tertiary care center
DOI:10.1086/432939
URL
PMID:16142662
[本文引用: 1]

BACKGROUND: During the past decade, Candida glabrata has emerged as an important cause of fungemia. We reviewed demographic data, risk factors, treatment, and outcomes associated with C. glabrata fungemia from 1995-2002 and performed susceptibility testing of isolates. METHODS: Data on all episodes of fungemia were prospectively recorded, and the associated isolates were saved. Medical records were reviewed retrospectively. Susceptibility testing was performed for fluconazole, itraconazole, and voriconazole. RESULTS: C. glabrata caused 103 (17%) of 609 fungemic episodes during the 8-year period that we studied. Medical records and isolates were available for 94 episodes that occurred in 91 patients. The patients included 42 men and 49 women. The mean age was 51 years. Thirty-four episodes (36%) occurred in patients >60 years old; only 3 episodes occurred in patients <1 year old. The most common predisposing factors were use of broad-spectrum antibiotics (in 86% of episodes), use of central venous catheters (77%), stay in an intensive care unit (48%), renal failure (46%), and receipt of parenteral nutrition (45%). Of the 94 episodes, 83 were treated with antifungal agents. The overall mortality rate at day 30 was 29%. For the 11 episodes that were not treated, the mortality rate was 64% (7 of 11 episodes). Outcome appeared to be unrelated to whether fluconazole or amphotericin B was administered. In vitro, 60% of isolates were resistant to fluconazole, 83% to itraconazole, and 44% to voriconazole. Susceptibility to these azoles did not change over the 8 years of the study. CONCLUSION: C. glabrata fungemia was most often seen in older adults and was associated with a mortality rate of 29%. Outcomes appeared to be unrelated to in vitro susceptibility results and to the antifungal agent used.
Candida glabrata metallothioneins: cloning and sequence of the genes and characterization of proteins
Selective and tandem amplifification of a member of the metallothionein gene family in Candida glabrata
Inhibiting fungal multidrug resistance by disrupting an activator-mediator interaction
DOI:10.1038/nature16963
URL
PMID:26886795
[本文引用: 1]

Eukaryotic transcription activators stimulate the expression of specific sets of target genes through recruitment of co-activators such as the RNA polymerase II-interacting Mediator complex. Aberrant function of transcription activators has been implicated in several diseases. However, therapeutic targeting efforts have been hampered by a lack of detailed molecular knowledge of the mechanisms of gene activation by disease-associated transcription activators. We previously identified an activator-targeted three-helix bundle KIX domain in the human MED15 Mediator subunit that is structurally conserved in Gal11/Med15 Mediator subunits in fungi. The Gal11/Med15 KIX domain engages pleiotropic drug resistance transcription factor (Pdr1) orthologues, which are key regulators of the multidrug resistance pathway in Saccharomyces cerevisiae and in the clinically important human pathogen Candida glabrata. The prevalence of C. glabrata is rising, partly owing to its low intrinsic susceptibility to azoles, the most widely used antifungal agent. Drug-resistant clinical isolates of C. glabrata most commonly contain point mutations in Pdr1 that render it constitutively active, suggesting that this transcriptional activation pathway represents a linchpin in C. glabrata multidrug resistance. Here we perform sequential biochemical and in vivo high-throughput screens to identify small-molecule inhibitors of the interaction of the C. glabrata Pdr1 activation domain with the C. glabrata Gal11A KIX domain. The lead compound (iKIX1) inhibits Pdr1-dependent gene activation and re-sensitizes drug-resistant C. glabrata to azole antifungals in vitro and in animal models for disseminated and urinary tract C. glabrata infection. Determining the NMR structure of the C. glabrata Gal11A KIX domain provides a detailed understanding of the molecular mechanism of Pdr1 gene activation and multidrug resistance inhibition by iKIX1. We have demonstrated the feasibility of small-molecule targeting of a transcription factor-binding site in Mediator as a novel therapeutic strategy in fungal infectious disease.
Prevalence, susceptibility profile and proteinase production of yeasts causing vulvovaginitis in Turkish women
Nosocomial fungal infections: epidemiology, diagnosis, and treatment
DOI:10.1080/13693780701218689
URL
PMID:17510856
[本文引用: 1]

Invasive fungal infections are increasingly common in the nosocomial setting. Furthermore, because risk factors for these infections continue to increase in frequency, it is likely that nosocomial fungal infections will continue to increase in frequency in the coming decades. The predominant nosocomial fungal pathogens include Candida spp., Aspergillus spp., Mucorales, Fusarium spp., and other molds, including Scedosporium spp. These infections are difficult to diagnose and cause high morbidity and mortality despite antifungal therapy. Early initiation of effective antifungal therapy and reversal of underlying host defects remain the cornerstones of treatment for nosocomial fungal infections. In recent years, new antifungal agents have become available, resulting in a change in standard of care for many of these infections. Nevertheless, the mortality of nosocomial fungal infections remains high, and new therapeutic and preventative strategies are needed.
Twenty years of the SENTRY antifungal surveillance program: results for Candida species from 1997-2016
Geographic variation in the susceptibilities of invasive isolates of Candida glabrata to seven systemically active antifungal agents: a global assessment from the ARTEMIS Antifungal Surveillance Program conducted in 2001 and 2002
DOI:10.1128/JCM.42.7.3142-3146.2004
URL
PMID:15243073
[本文引用: 1]

We examined the susceptibilities to amphotericin B, flucytosine, fluconazole, posaconazole, ravuconazole, voriconazole, and caspofungin of 601 invasive isolates of Candida glabrata and grouped the isolates by geographic location: North America (331 isolates), Latin America (58 isolates), Europe (135 isolates), and Asia-Pacific (77 isolates). Caspofungin (MIC at which 90% of isolates tested are susceptible [MIC(90)], 0.12 microg/ml; 100% of strains are susceptible [S] at a MIC of
Formation of new chromosomes as a virulence mechanism in yeast Candida glabrata
Fungal biofilm resistance
Aspartyl proteases in Candida glabrata are required for suppression of the host innate immune response
Role of ATP-binding- cassette transporter genes in high-frequency acquisition of resistance to azole antifungals in Candida glabrata
DOI:10.1128/AAC.45.4.1174-1183.2001
URL
PMID:11257032
[本文引用: 1]

Candida glabrata has been often isolated from AIDS patients with oropharyngeal candidiasis treated with azole antifungal agents, especially fluconazole. We recently showed that the ATP-binding-cassette (ABC) transporter gene CgCDR1 was upregulated in C. glabrata clinical isolates resistant to azole antifungal agents (D. Sanglard, F. Ischer, D. Calabrese, P. A. Majcherczyk, and J. Bille, Antimicrob. Agents Chemother. 43:2753-2765, 1999). Deletion of CgCDR1 in C. glabrata rendered the null mutant hypersusceptible to azole derivatives and showed the importance of this gene in mediating azole resistance. We observed that wild-type C. glabrata exposed to fluconazole in a medium containing the drug at 50 microg/ml developed resistance to this agent and other azoles at a surprisingly high frequency (2 x 10(-4) to 4 x 10(-4)). We show here that this high-frequency azole resistance (HFAR) acquired in vitro was due, at least in part, to the upregulation of CgCDR1. The CgCDR1 deletion mutant DSY1041 could still develop HFAR but in a medium containing fluconazole at 5 microg/ml. In the HFAR strain derived from DSY1041, a distinct ABC transporter gene similar to CgCDR1, called CgCDR2, was upregulated. This gene was slightly expressed in clinical isolates but was upregulated in strains with the HFAR phenotype. Deletion of both CgCDR1 and CgCDR2 suppressed the development of HFAR in a medium containing fluconazole at 5 microg/ml, showing that both genes are important mediators of resistance to azole derivatives in C. glabrata. We also show here that the HFAR phenomenon was linked to the loss of mitochondria in C. glabrata. Mitochondrial loss could be obtained by treatment with ethidium bromide and resulted in acquisition of resistance to azole derivatives without previous exposure to these agents. Azole resistance obtained in vitro by HFAR or by agents stimulating mitochondrial loss was at least linked to the upregulation of both CgCDR1 and CgCDR2.
Interaction of human fibronectin with Candida glabrata epithelial adhesin 6 (Epa6)
DOI:10.18388/abp.2016_1328
URL
PMID:27474404
[本文引用: 1]

Adherence of pathogens to extracellular matrix proteins and host cells is one of the essential steps in the microbial colonization of the human organism. The adhesion of C. glabrata, i.e. the second major causative agent of human disseminated candidiases after C. albicans, to the host epithelium mainly engages specific fungal cell wall proteins - epithelial adhesins (Epa) - in particular, Epa1, Epa6 and Epa7. The aim of the present study was to identify the major Epa protein involved in the interactions with the human extracellular matrix protein - fibronectin - and to present the kinetic and thermodynamic characteristics of these interactions. A relatively novel gel-free approach, i.e. the
Biofilms of non-Candida albicans Candida species: quantification, structure and matrix composition
In vitro biofilm activity of non-Candida albicans Candida species
DOI:10.1007/s00284-010-9649-7
URL
PMID:20401483
[本文引用: 1]

Candidosis has been attributed to C. albicans; however, infections caused by non-Candida albicans Candida (NCAC) species are increasingly being recognised. The ability of Candida to grow as a biofilm is an important feature that promotes both infection and persistence in the host. The biofilms' activity is significant since high activity might be associated with enhanced expression of putative virulence factors, whilst in contrast low activity has previously been suggested as a mechanism for resistance of biofilm cells to antimicrobials. The aim of this study was to determine the metabolic activity of in vitro biofilms formed by different clinical isolates of NCAC species. The in situ total metabolic activity of C. parapsilosis, C. tropicalis and C. glabrata biofilms was determined using 2,3-(2-methoxy-4-nitro-5-sulphophenyl)-5-[(phenylamino) carbonyl]-2H-tetrazolium hydroxide (XTT) reduction assay, and the number of cultivable cells was also established by CFU (colony forming unit) counts. The biofilm structure was assessed by scanning electron microscopy (SEM). Results showed that total biofilm metabolic activity was species and strain dependent. C. glabrata exhibited the lowest biofilm metabolic activity despite having the highest number of biofilm cultivable cells. Similarly, the metabolic activity of resuspended C. glabrata biofilm and planktonic cells was lower than that of the other species. This study demonstrates the existence of intrinsic activity differences amongst NCAC species, which could have important implications in terms of species relative virulence. Furthermore, the absence of an obvious correlation, between cultivable cells number and total biofilm activity, raises the question about which parameter is the most appropriate for the in vitro assessment of biofilms and their potential clinical significance.
Adherence and biofilm formation of non-Candida albicans Candida species
DOI:10.1016/j.tim.2011.02.003
URL
PMID:21411325
[本文引用: 1]

Most cases of candidosis have been attributed to Candida albicans, but recently non-C. albicans Candida species have been identified as frequent human pathogens. Candida pathogenicity has been attributed to several factors, including adhesion to medical devices and/or host cells, biofilm formation, and secretion of hydrolytic enzymes (proteases, phospholipases and haemolysins). Although 'new' Candida species are emerging, there is still a lack of information about their pathogenicity. This review discusses recent advances in our knowledge of Candida glabrata, Candida parapsilosis and Candida tropicalis virulence factors, specifically those of adhesion and biofilm formation, which are key components in Candida pathogenicity.
Candida glabrata, Candida parapsilosis and Candida tropicalis: biology, epidemiology, pathogenicity and antifungal resistance
DOI:10.1111/j.1574-6976.2011.00278.x
URL
PMID:21569057
[本文引用: 1]

The incidence of infections caused by Candida species (candidosis) has increased considerably over the past three decades, mainly due to the rise of the AIDS epidemic, an increasingly aged population, higher numbers of immunocompromised patients and the more widespread use of indwelling medical devices. Candida albicans is the main cause of candidosis; however, non-C. albicans Candida (NCAC) species such as Candida glabrata, Candida tropicalis and Candida parapsilosis are now frequently identified as human pathogens. The apparent increased emergence of these species as human pathogens can be attributed to improved identification methods and also associated with the degree of diseases of the patients, the interventions that they were subjected and the drugs used. Candida pathogenicity is facilitated by a number of virulence factors, most importantly adherence to host surfaces including medical devices, biofilm formation and secretion of hydrolytic enzymes (e.g. proteases, phospholipases and haemolysins). Furthermore, despite extensive research to identify pathogenic factors in fungi, particularly in C. albicans, relatively little is known about NCAC species. This review provides information on the current state of knowledge on the biology, identification, epidemiology, pathogenicity and antifungal resistance of C. glabrata, C. parapsilosis and C. tropicalis.
Dark brown is the more virulent of the switch phenotypes of Candida glabrata
DOI:10.1099/mic.0.2008/020578-0 URL [本文引用: 1]
Coevolution of morphology and virulence in Candida species
DOI:10.1128/EC.05085-11
URL
PMID:21764907
[本文引用: 1]

Many of the major human fungal pathogens are known to undergo morphological changes, which in certain cases are associated with virulence. Although there has been an intense research focus on morphology in fungi, very little is known about how morphology evolved in conjunction with a variety of other virulence properties. However, several recent important discoveries, primarily in Candida species, are beginning to shed light on this important area and answer many longstanding questions. In this minireview, we first provide a description of the major fungal morphologies, as well as the roles of morphology and morphology-associated gene expression in virulence. Next, focusing largely on Candida species, we examine the evolutionary relationships among specific morphological forms. Finally, drawing on recent findings, we begin to address the question of how specific morphological changes came to be associated with virulence of Candida species during evolution.
Adhesins in Candida glabrata
Gain-of-function mutations in PDR1, a regulator of antifungal drug resistance in Candida glabrata, control adherence to host cells
DOI:10.1128/IAI.00074-13
URL
PMID:23460523
[本文引用: 1]

Candida glabrata is an emerging opportunistic pathogen that is known to develop resistance to azole drugs due to increased drug efflux. The mechanism consists of CgPDR1-mediated upregulation of ATP-binding cassette transporters. A range of gain-of-function (GOF) mutations in CgPDR1 have been found to lead not only to azole resistance but also to enhanced virulence. This implicates CgPDR1 in the regulation of the interaction of C. glabrata with the host. To identify specific CgPDR1-regulated steps of the host-pathogen interaction, we investigated in this work the interaction of selected CgPDR1 GOF mutants with murine bone marrow-derived macrophages and human acute monocytic leukemia cell line (THP-1)-derived macrophages, as well as different epithelial cell lines. GOF mutations in CgPDR1 did not influence survival and replication within macrophages following phagocytosis but led to decreased adherence to and uptake by macrophages. This may allow evasion from the host's innate cellular immune response. The interaction with epithelial cells revealed an opposite trend, suggesting that GOF mutations in CgPDR1 may favor epithelial colonization of the host by C. glabrata through increased adherence to epithelial cell layers. These data reveal that GOF mutations in CgPDR1 modulate the interaction with host cells in ways that may contribute to increased virulence.
Upregulation of the adhesin gene EPA1 mediated by PDR1 in Candida glabrata leads to enhanced host colonization
DOI:10.1128/mSphere.00065-15
URL
PMID:27303714
[本文引用: 1]

Candida glabrata is the second most common Candida species causing disseminated infection, after C. albicans. C. glabrata is intrinsically less susceptible to the widely used azole antifungal drugs and quickly develops secondary resistance. Resistance typically relies on drug efflux with transporters regulated by the transcription factor Pdr1. Gain-of-function (GOF) mutations in PDR1 lead to a hyperactive state and thus efflux transporter upregulation. Our laboratory has characterized a collection of C. glabrata clinical isolates in which azole resistance was found to correlate with increased virulence in vivo. Contributing phenotypes were the evasion of adhesion and phagocytosis by macrophages and an increased adhesion to epithelial cells. These phenotypes were found to be dependent on PDR1 GOF mutation and/or C. glabrata strain background. In the search for the molecular effectors, we found that PDR1 hyperactivity leads to overexpression of specific cell wall adhesins of C. glabrata. Further study revealed that EPA1 regulation, in particular, explained the increase in adherence to epithelial cells. Deleting EPA1 eliminates the increase in adherence in an in vitro model of interaction with epithelial cells. In a murine model of urinary tract infection, PDR1 hyperactivity conferred increased ability to colonize the bladder and kidneys in an EPA1-dependent way. In conclusion, this study establishes a relationship between PDR1 and the regulation of cell wall adhesins, an important virulence attribute of C. glabrata. Furthermore, our data show that PDR1 hyperactivity mediates increased adherence to host epithelial tissues both in vitro and in vivo through upregulation of the adhesin gene EPA1. IMPORTANCE Candida glabrata is an important fungal pathogen in human diseases and is also rapidly acquiring drug resistance. Drug resistance can be mediated by the transcriptional activator PDR1, and this results in the upregulation of multidrug transporters. Intriguingly, this resistance mechanism is associated in C. glabrata with increased virulence in animal models and also with increased adherence to specific host cell types. The C. glabrata adhesin gene EPA1 is a major contributor of virulence and adherence to host cells. Here, we show that EPA1 expression is controlled by PDR1 independently of subtelomeric silencing, a known EPA1 regulation mechanism. Thus, a relationship exists between PDR1, EPA1 expression, and adherence to host cells, which is critical for efficient virulence. Our results demonstrate that acquisition of drug resistance is beneficial for C. glabrata in fungus-host relationships. These findings further highlight the challenges of the therapeutic management of C. glabrata infections in human patients.
A nonsense mutation in the ERG6 gene leads to reduced susceptibility to polyenes in a clinical isolate of Candida glabrata
DOI:10.1128/AAC.00423-08
URL
PMID:18694952
[本文引用: 1]

Unlike the molecular mechanisms that lead to azole drug resistance, the molecular mechanisms that lead to polyene resistance are poorly documented, especially in pathogenic yeasts. We investigated the molecular mechanisms responsible for the reduced susceptibility to polyenes of a clinical isolate of Candida glabrata. Sterol content was analyzed by gas-phase chromatography, and we determined the sequences and levels of expression of several genes involved in ergosterol biosynthesis. We also investigated the effects of the mutation harbored by this isolate on the morphology and ultrastructure of the cell, cell viability, and vitality and susceptibility to cell wall-perturbing agents. The isolate had a lower ergosterol content in its membranes than the wild type, and the lower ergosterol content was found to be associated with a nonsense mutation in the ERG6 gene and induction of the ergosterol biosynthesis pathway. Modifications of the cell wall were also seen, accompanied by increased susceptibility to cell wall-perturbing agents. Finally, this mutation, which resulted in a marked fitness cost, was associated with a higher rate of cell mortality. Wild-type properties were restored by complementation of the isolate with a centromeric plasmid containing a wild-type copy of the ERG6 gene. In conclusion, we have identified the molecular event responsible for decreased susceptibility to polyenes in a clinical isolate of C. glabrata. The nonsense mutation detected in the ERG6 gene of this isolate led to a decrease in ergosterol content. This isolate may constitute a useful tool for analysis of the relevance of protein trafficking in the phenomena of azole resistance and pseudohyphal growth.
From attachment to damage: defined genes of Candida albicans mediate adhesion, invasion and damage during interaction with oral epithelial cells
DOI:10.1371/journal.pone.0017046
URL
PMID:21407800
[本文引用: 1]

Candida albicans frequently causes superficial infections by invading and damaging epithelial cells, but may also cause systemic infections by penetrating through epithelial barriers. C. albicans is an unusual pathogen because it can invade epithelial cells via two distinct mechanisms: induced endocytosis, analogous to facultative intracellular enteropathogenic bacteria, and active penetration, similar to plant pathogenic fungi. Here we investigated the molecular basis of C. albicans epithelial interactions. By systematically assessing the contributions of defined fungal pathways and factors to different stages of epithelial interactions, we provide an expansive portrait of the processes and activities involved in epithelial infection. We strengthen the concept that hyphal formation is critical for epithelial invasion. Importantly, our data support a model whereby initial epithelial invasion per se does not elicit host damage, but that C. albicans relies on a combination of contact-sensing, directed hyphal extension, active penetration and the expression of novel pathogenicity factors for further inter-epithelial invasion, dissemination and ultimate damage of host cells. Finally, we explore the transcriptional landscape of C. albicans during the early stages of epithelial interaction, and, via genetic analysis, identify ICL1 and PGA34 as novel oral epithelial pathogenicity factors.
Candida glabrata and Candida albicans; dissimilar tissue tropism and infectivity in a gnotobiotic model of mucosal candidiasis
DOI:10.1111/j.1574-695X.2007.00287.x
URL
PMID:17854475
[本文引用: 2]

Germ-free transgenic epsilon 26 (Tgepsilon26) mice, deficient in both natural killer (NK)- and T-cells, were inoculated (orally) with each of two Candida glabrata (BG2 or BG1003) or Candida albicans (CAF2-1 or SC5314) strains. Candida glabrata- or C. albicans-colonized mice exhibited similar numbers of viable Candida in the alimentary tract. Neither C. glabrata nor C. albicans caused systemic candidiasis of endogenous (alimentary tract) origin. Candida albicans invaded oroesophageal (tongue, palate, esophagus) and keratinized gastric tissues, evoked hyperkeratosis and a prominent, chronic, granulocyte-dominated, inflammatory response in all infected tissues, stimulated the production of splenic granulocytes and was lethal for the mice within 3-5 weeks after oral colonization. The two C. glabrata strains colonized the alimentary tract and penetrated into the keratinized (cardia-antrum) gastric tissues, but in contrast to C. albicans, were unable to infect oroesophageal tissues. Furthermore, C. glabrata strains were not lethal for the Tgepsilon26 mice, and did not evoke an inflammatory response in colonized gastric tissues or stimulate the production of splenic granulocytes. This 'stealth-like' behavior could explain the ability of C. glabrata to persist in infected tissues and survive as a commensal in the alimentary tract.
Antifungal resistance: current trends and future strategies to combat
DOI:10.2147/IDR.S124918
URL
PMID:28919789
[本文引用: 1]

Antifungal resistance represents a major clinical challenge to clinicians responsible for treating invasive fungal infections due to the limited arsenal of systemically available antifungal agents. In addition current drugs may be limited by drug-drug interactions and serious adverse effects/toxicities that prevent their prolonged use or dosage escalation. Fluconazole resistance is of particular concern in non-Candida albicans species due to the increased incidence of infections caused by these species in different geographic locations worldwide and the elevated prevalence of resistance to this commonly used azole in many institutions. C. glabrata resistance to the echinocandins has also been documented to be rising in several US institutions, and a higher percentage of these isolates may also be azole resistant. Azole resistance in Aspergillus fumigatus due to clinical and environmental exposure to this class of agents has also been found worldwide, and these isolates can cause invasive infections with high mortality rates. In addition, several species of Aspergillus, and other molds, including Scedosporium and Fusarium species, have reduced susceptibility or pan-resistance to clinically available antifungals. Various investigational antifungals are currently in preclinical or clinical development, including several of them that have the potential to overcome resistance observed against the azoles and the echinocandins. These include agents that also target ergosterol and b-glucan biosynthesis, as well as compounds with novel mechanisms of action that may also overcome the limitations of currently available antifungal classes, including both resistance and adverse effects/toxicity.
Molecular evidence for an ancient duplication of the entire yeast genome
DOI:10.1038/42711
URL
PMID:9192896
[本文引用: 3]

Gene duplication is an important source of evolutionary novelty. Most duplications are of just a single gene, but Ohno proposed that whole-genome duplication (polyploidy) is an important evolutionary mechanism. Many duplicate genes have been found in Saccharomyces cerevisiae, and these often seem to be phenotypically redundant. Here we show that the arrangement of duplicated genes in the S. cerevisiae genome is consistent with Ohno's hypothesis. We propose a model in which this species is a degenerate tetraploid resulting from a whole-genome duplication that occurred after the divergence of Saccharomyces from Kluyveromyces. Only a small fraction of the genes were subsequently retained in duplicate (most were deleted), and gene order was rearranged by many reciprocal translocations between chromosomes. Protein pairs derived from this duplication event make up 13% of all yeast proteins, and include pairs of transcription factors, protein kinases, myosins, cyclins and pheromones. Tetraploidy may have facilitated the evolution of anaerobic fermentation in Saccharomyces.